Abstract
Additive models and polygenic risk frameworks have become standard approaches in human genetics, aggregating variant effects into composite estimates of phenotype. While effective in certain contexts, these models do not fully capture the structured, interactive nature of biological systems.
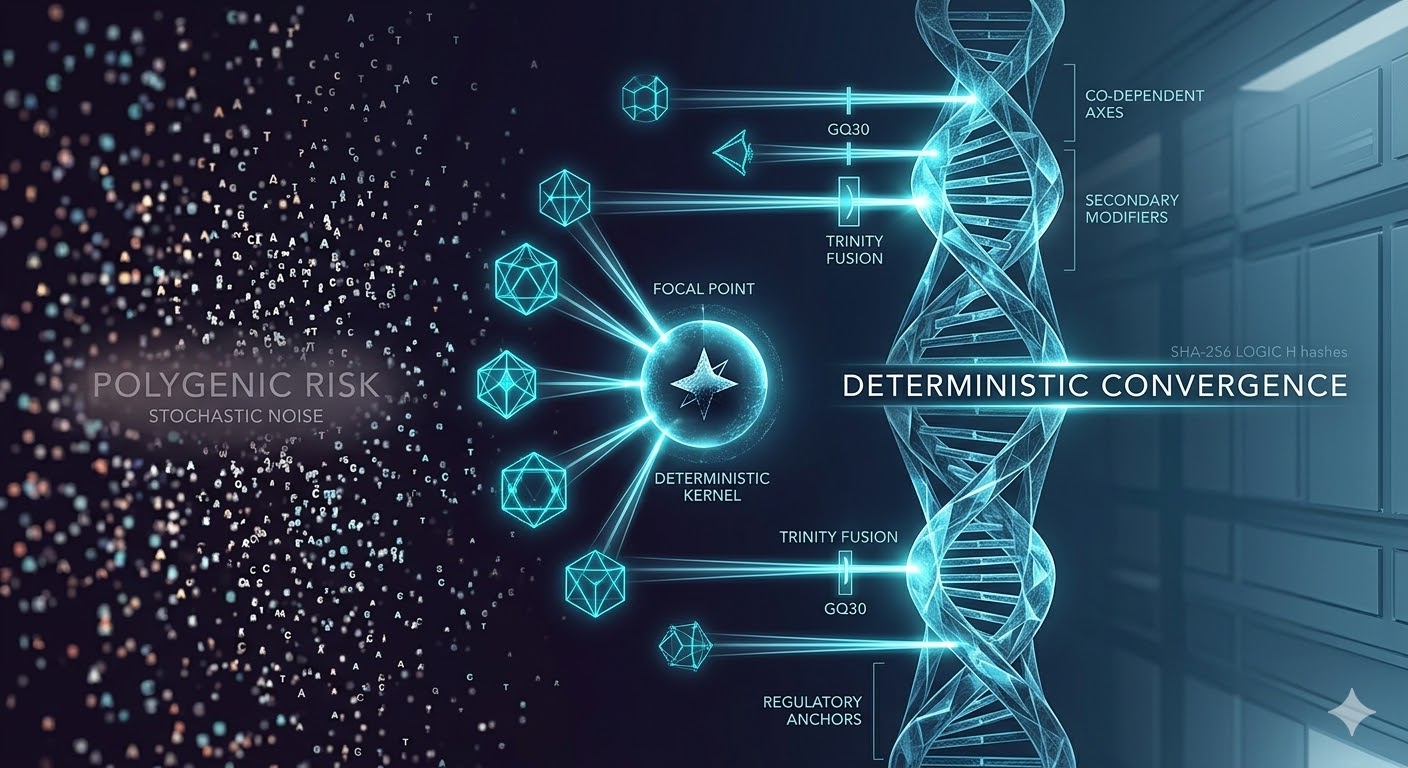
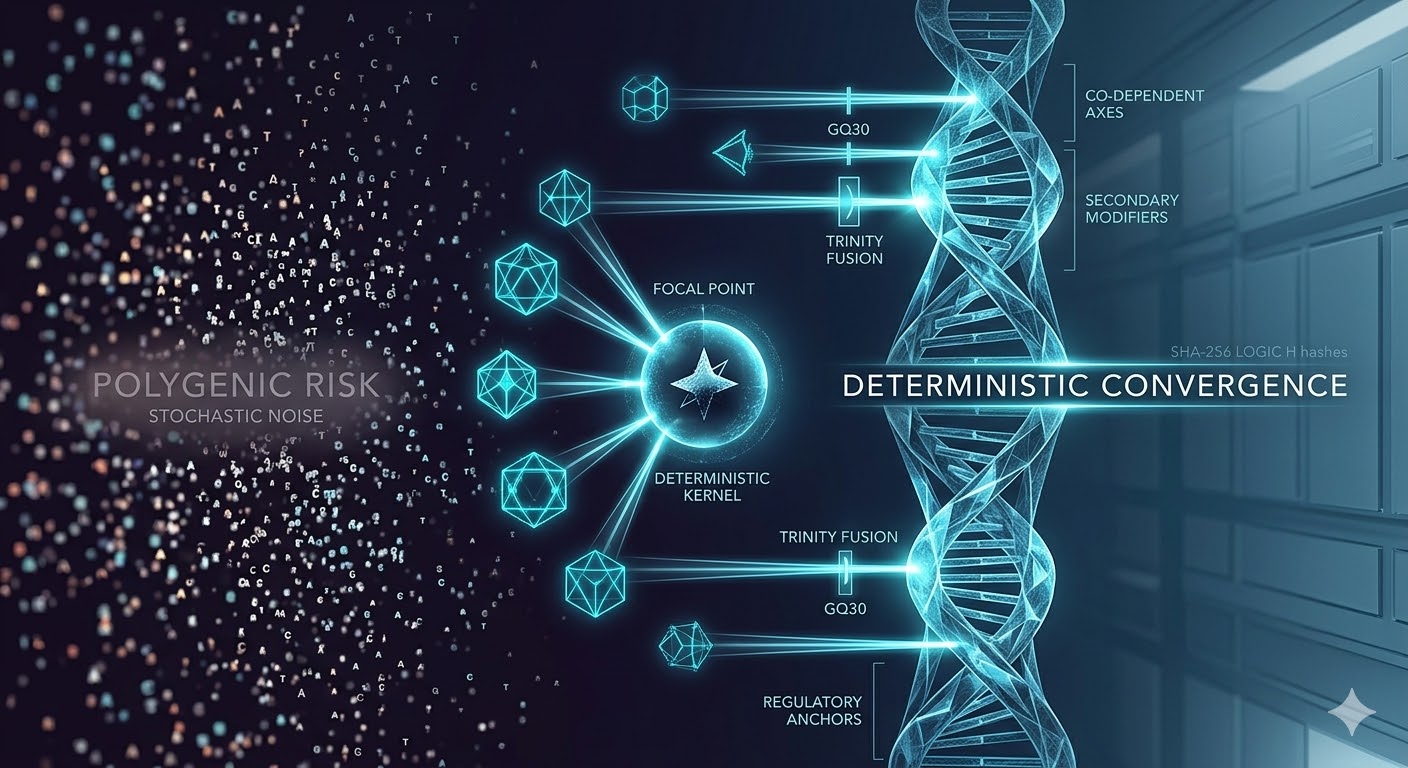
Here, we introduce a computational framework that evaluates genomic systems through deterministic convergence—the repeated emergence of stable multi-variant configurations across independent simulation runs under identical conditions. Using Proteus V3, we observe that genomic systems frequently resolve into low-dimensional hierarchies, with some phenotypes reducible to dominant genetic axes and others dependent on co-equal, distributed interactions.
Across cardiovascular, oncological, metabolic, and neurological domains, we find that many high-complexity variant combinations fail to persist under validation, while a subset of configurations consistently converges and remains stable under perturbation. These results suggest that genomic architecture may be better understood as structured and constrained rather than diffuse and additive.
This framework does not establish causality but provides a reproducible method for distinguishing stable biological signal from stochastic variability. If validated across larger cohorts, deterministic convergence may offer a foundation for reinterpreting genetic architecture as a problem of structure rather than aggregation.
Introduction
The prevailing computational paradigm in human genetics assumes that variants contribute additively and, to a first approximation, independently to phenotype. Polygenic risk score (PRS) frameworks extend this logic by aggregating many small effects into a composite estimate of risk. Although these approaches have demonstrated utility, they do not fully reflect the organizational principles of biological systems.
Biological processes are inherently interactive. Proteins function within networks, regulatory pathways impose constraints, and cellular behavior emerges from coordinated systems rather than isolated loci. Under these conditions, an additive approximation may obscure structure rather than reveal it.
If biological systems are structured, then genomic interpretation requires models capable of identifying interaction hierarchies rather than simply accumulating effects. The central question becomes not how much signal is present, but how that signal is organized.
Conceptual Framework: Deterministic Convergence
A central concept emerging from this work is deterministic convergence.
Deterministic convergence refers to the repeated emergence of the same multi-variant configurations across independent simulation runs under identical initial conditions. In this framework, convergence is not assumed but measured.
It is quantified through:
concordance of variant composition across runs
preservation of rank-order within retained configurations
stability of relative fitness under perturbation and resampling
If independent runs produce different dominant configurations under identical conditions, the system is sampling variability rather than modeling constraint.
Conversely, when the same configurations repeatedly emerge and persist under validation pressure, this indicates the presence of underlying constraint rather than stochastic signal.
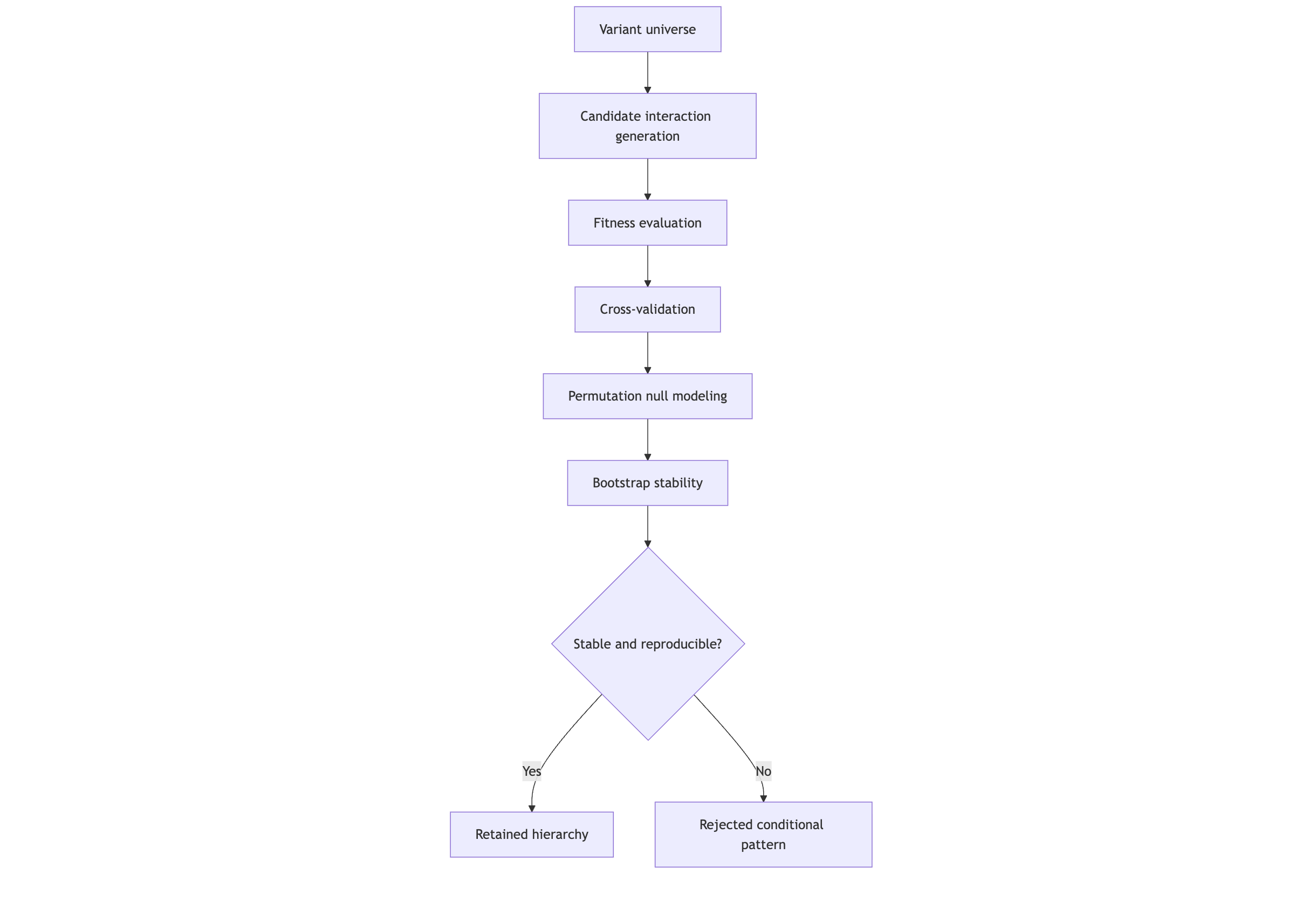
Methods Overview
Proteus V3 was developed to explore genomic structure under conditions that enforce reproducibility and validation.
Rather than evaluating variants in isolation, the framework generates candidate multi-variant configurations and subjects them to iterative validation pressure, including:
cross-validation across subsampled cohorts
permutation-based null modeling
bootstrap resampling for stability assessment
Only configurations that demonstrate deterministic convergence—remaining stable, reproducible, and interpretable across independent runs—are retained. Configurations that fail to persist are rejected as unstable or non-informative.
The internal scoring functions and optimization mechanisms are proprietary and not disclosed in this work.
Results
Deterministic convergence reveals hierarchical organization
Across multiple biological domains, a consistent pattern of deterministic convergence emerged: high-dimensional genomic space frequently resolves into low-dimensional structure.
In cardiovascular analyses, simulations repeatedly converged on a dominant structural axis, with a smaller modifier layer contributing secondary effects. Oncological analyses similarly converged on a primary regulatory anchor, with additional variants providing limited incremental contribution.
These systems appear reducible: although many variants are present, a small subset defines the core structure of the phenotype.
Distributed systems do not collapse to a single axis
In contrast, neurological analyses demonstrated a different form of convergence. Rather than collapsing to a single dominant configuration, multiple co-dependent variant sets persisted together across runs.
These systems exhibit distributed architecture, requiring simultaneous integrity across multiple biological axes, including protein handling, transcriptional regulation, and degradation pathways.
This distinction suggests that convergence behavior itself can be used to classify genomic systems as:
reducible (dominant-axis driven)
distributed (multi-axis dependent)
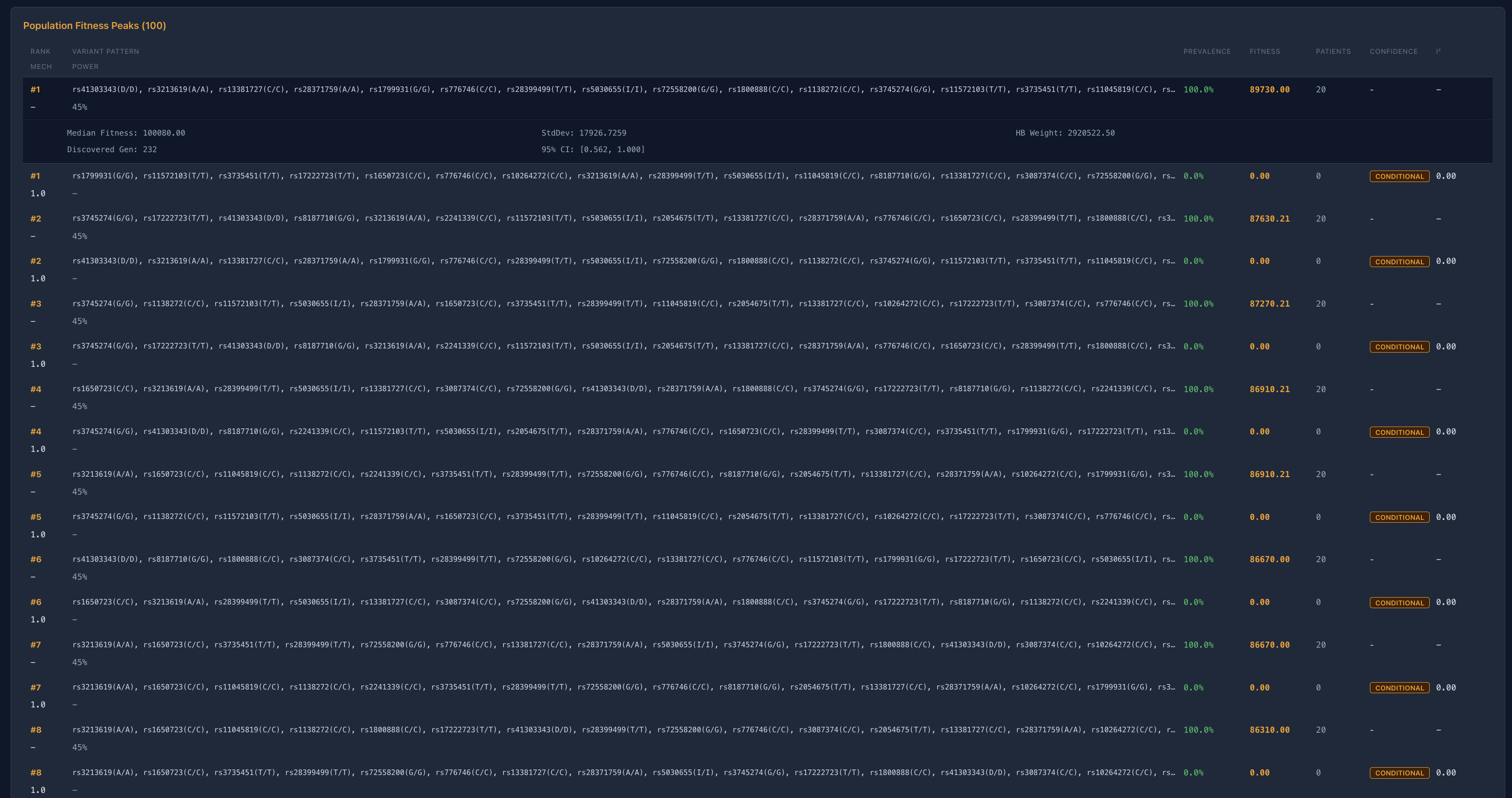
Rejection of unstable combinatorial space
A second notable observation is the systematic rejection of high-complexity variant combinations.
Across domains, many candidate configurations failed to exhibit deterministic convergence, often showing:
zero recurrence across runs
instability under perturbation
no consistent contribution to fitness
This suggests that a substantial portion of high-dimensional genomic space does not exhibit deterministic convergence and therefore may represent unstable or non-informative structure rather than durable biological signal.
Interpretation
These observations support a shift in how genomic systems are conceptualized.
Rather than viewing phenotype as the sum of independent effects, this framework suggests that phenotype may emerge from constrained interaction structures. Deterministic convergence provides a mechanism for distinguishing between:
stable, biologically meaningful configurations
transient or stochastic combinations
Importantly, this framework does not assume that all systems converge. In some cases, lack of convergence is itself an informative property, indicating distributed dependency rather than model failure.
A model that does not produce consistent convergence under identical conditions is sampling variability, not identifying constraint.
Limitations
These findings should be interpreted conservatively.
Current analyses are limited by:
cohort size and composition
incomplete population diversity
reliance on computational rather than experimental validation
The results do not establish causality and should not be interpreted as clinical evidence. Instead, they represent reproducible computational observations that generate testable hypotheses.
Implications
If validated across larger and more diverse datasets, deterministic convergence may provide a basis for reinterpreting genetic architecture.
This approach enables:
classification of genomic systems as reducible or distributed
identification of dominant versus secondary contributors
filtering of non-reproducible combinatorial noise
This represents a shift from aggregation to structure, with potential implications for:
target discovery
patient stratification
mechanistic interpretation of genetic variation
Conclusion
Genomic systems may be better understood as structured and constrained rather than purely additive.
By enforcing reproducibility through deterministic convergence, it becomes possible to distinguish stable biological signal from stochastic variability.
Proteus V3 provides a framework for testing this hypothesis. Whether these structures persist across larger and more diverse datasets remains an open and testable question.
Figures
Figure 1 — Hierarchical organization vs additivity
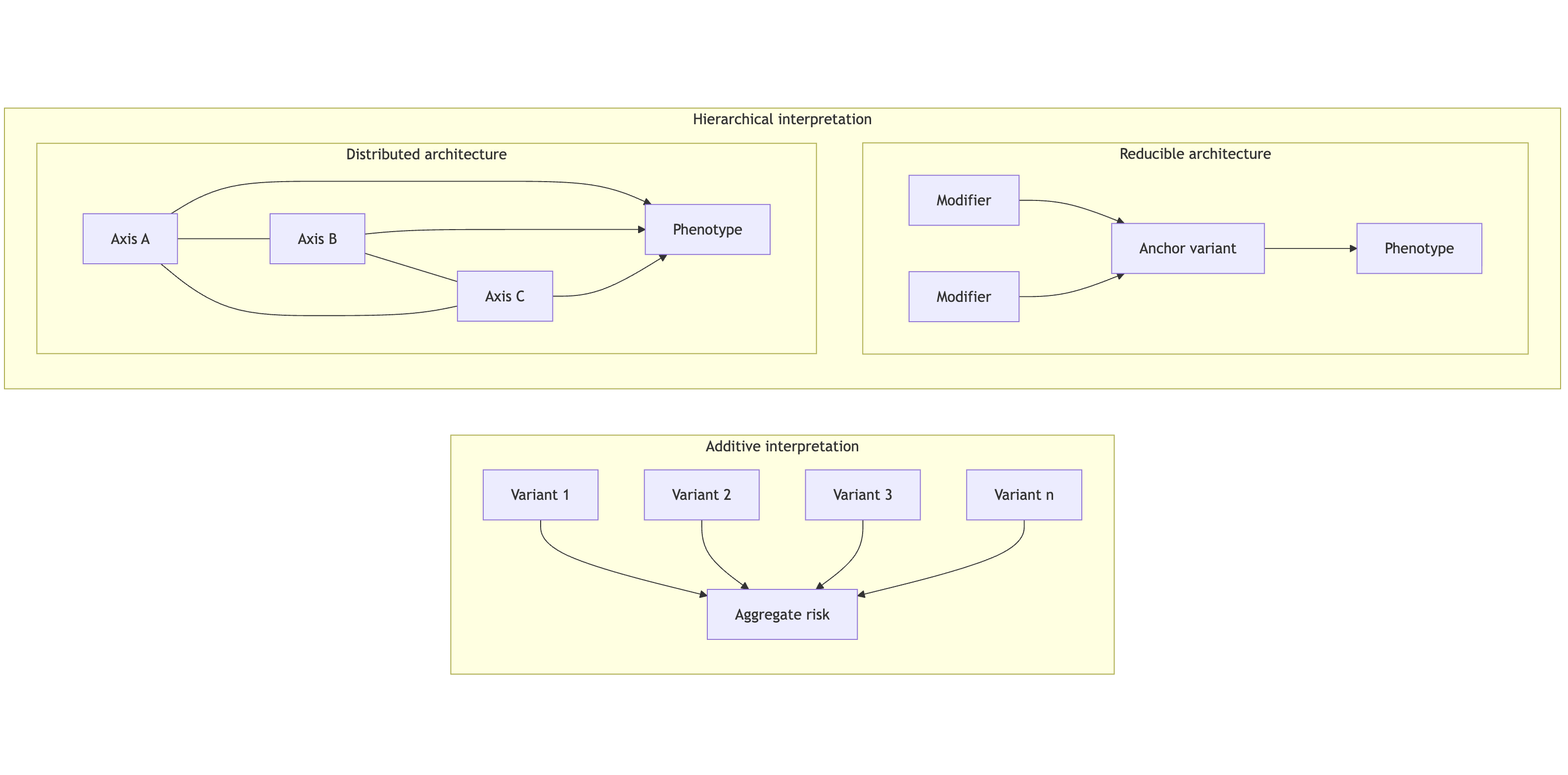
Figure 2 — Validation pipeline and convergence filtering