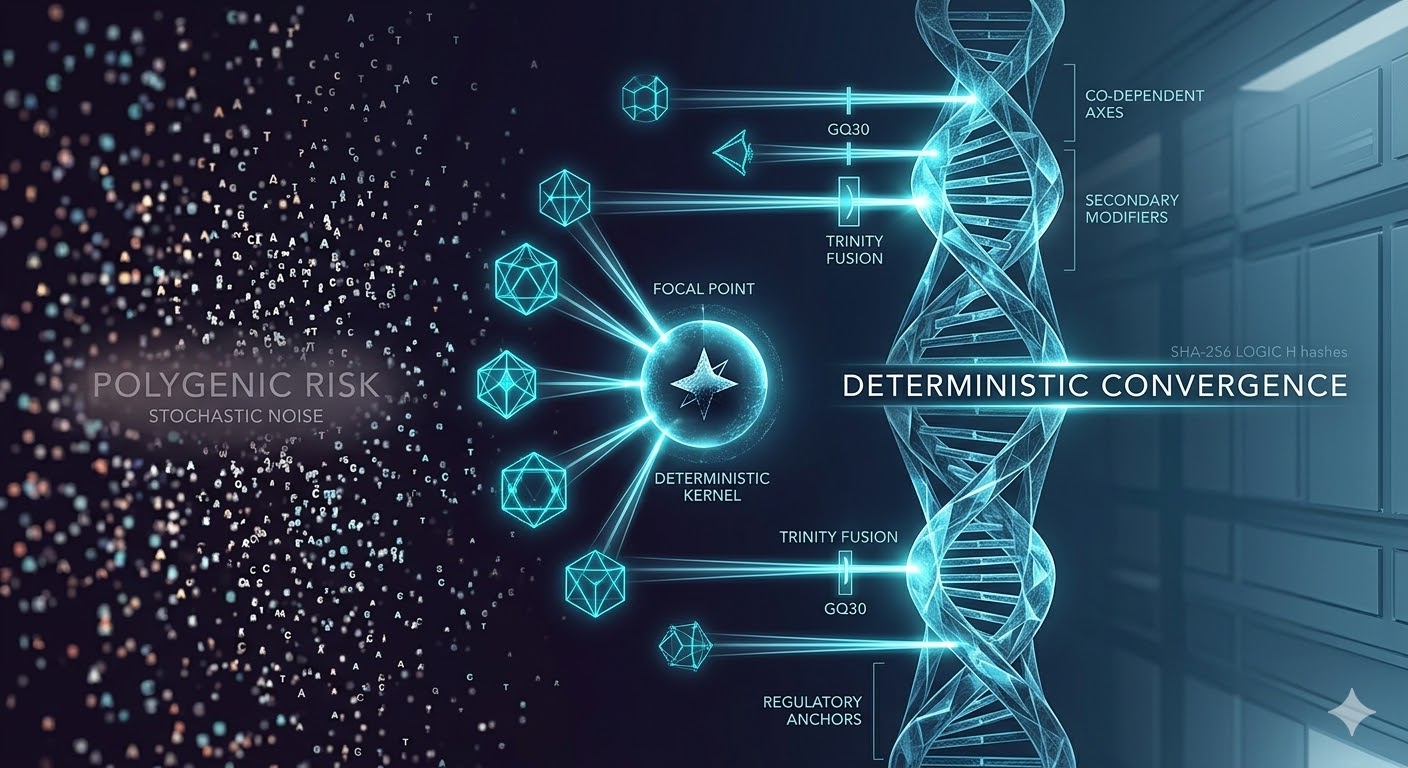
PROTEUS V3
Deterministic Convergence Engine
for Drug Discovery & Development
From static genomics to dynamic biological discovery.
NomosLogic, Inc.
EXECUTIVE OVERVIEW
Drug discovery is constrained by time, cost, and uncertainty — particularly in understanding how genetic variation drives drug response, resistance emergence, and late-stage trial failure. Current analytical frameworks treat genomic data as static datasets and variants as independent signals. Biology operates differently. It evolves, reconfigures, and adapts under pressure through interaction architectures that static analysis cannot detect.
PROTEUS V3 is a deterministic evolutionary simulation engine that models biological systems as dynamic processes. By simulating genomic interactions under selective pressure, PROTEUS V3 identifies deterministic convergence — the reproducible emergence of stable, multi-variant interaction structures across independent simulations under identical conditions. This enables earlier identification of resistance pathways, constrained interaction structures, and high-risk genomic configurations that drive treatment failure.
Unlike probabilistic AI/ML systems, PROTEUS V3 does not infer associations from data. It executes deterministic simulation against 17M+ validated biological and pharmacogenomic assets, producing outputs that are reproducible, traceable, and grounded in underlying biological constraint. Same inputs. Same outputs. Every time.
THE PROBLEM PROTEUS V3 ADDRESSES
Pharmaceutical development faces persistent structural challenges that current analytical approaches cannot resolve:
~90%
Oncology Drug Trial Failure Rate
The majority of oncology drugs entering clinical trials fail. A significant proportion fail not because the therapeutic mechanism is wrong, but because patient stratification and resistance prediction rely on single-variant association models that are structurally blind to multi-variant interaction architectures.
$2.6B
Average Cost to Bring a Drug to Market
Late-stage failures — driven by unforeseen resistance, patient variability, and incomplete understanding of genomic interactions — account for the majority of this cost. Reducing late-stage failure rates by even a small margin translates to billions in recovered development investment.
88.5%
Genomic Systems Exhibit Distributed Architecture
Across 26 independent PROTEUS studies spanning cardiovascular, neurological, oncological, and hepatic domains, 88.5% of observed biological systems operate as distributed constraint architectures. Variant influence is spread across interacting sets, not concentrated in single drivers. Single-target drug development strategies are structurally incomplete.
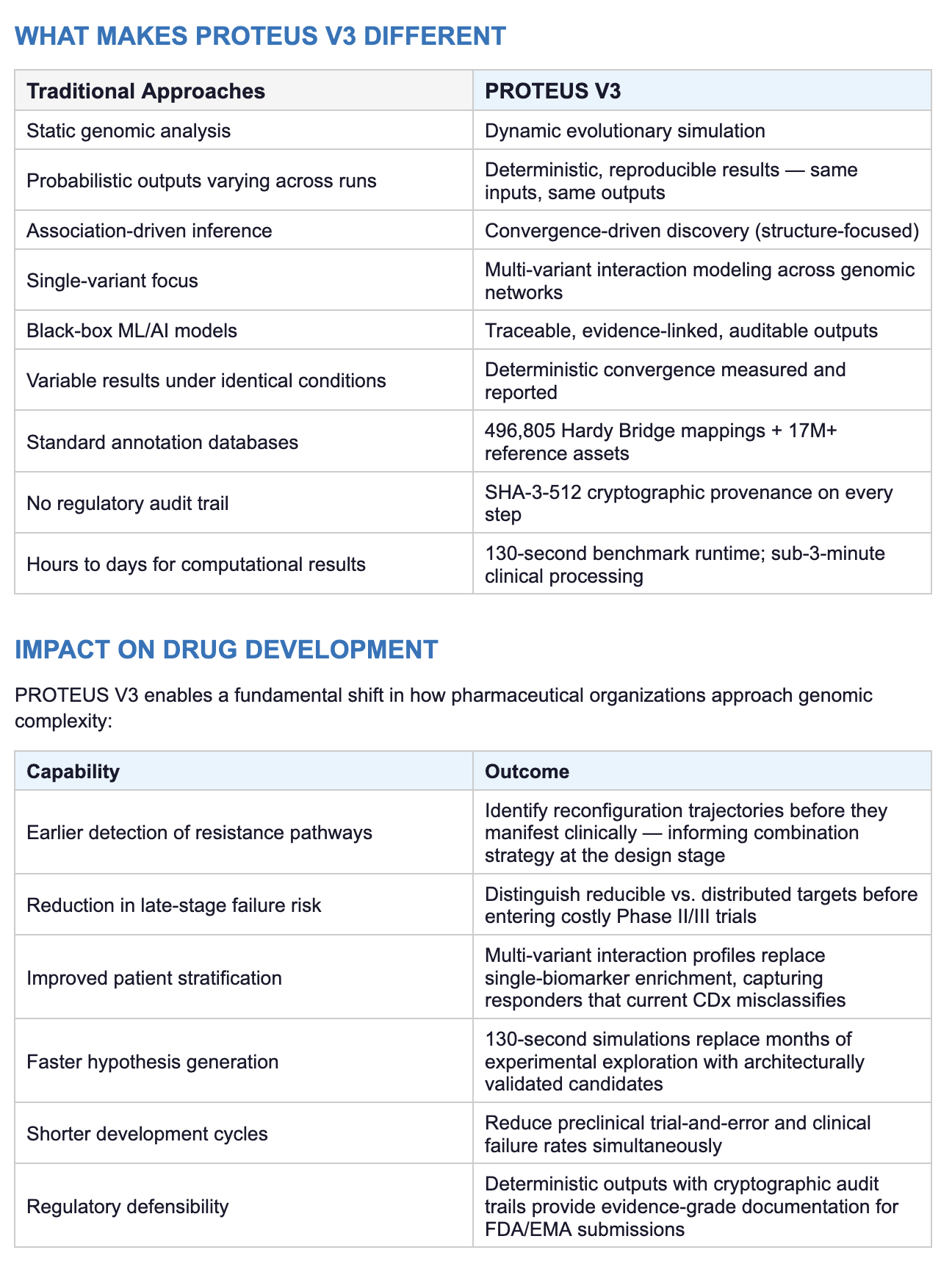
WHY CURRENT APPROACHES FALL SHORT
Existing genomic analytical frameworks — whether statistical, machine learning-based, or AI-driven — share fundamental limitations that PROTEUS V3 was designed to address:
Limitation
Consequence
PROTEUS V3 Response
Static analysis of dynamic systems
Identifies what is present but cannot model what will emerge, adapt, or reconfigure under therapeutic pressure
Evolutionary simulation models genomic change across generations under selective pressure, revealing trajectories invisible to static snapshots
Single-variant focus
Misses multi-variant interaction effects that drive treatment response, resistance, and patient variability
Multi-variant interaction modeling detects emergent behavior across genomic networks and constrained interaction structures
Probabilistic outputs
Different outputs across runs. Results cannot be reproduced, audited, or defended for regulatory submission
Deterministic execution: identical inputs produce identical outputs. SHA-3-512 cryptographic provenance on every analytical step
Association-driven inference
Identifies statistical correlations but not underlying biological structure. Cannot distinguish constraint from variability
Convergence-driven discovery: identifies constrained, reproducible interaction structures grounded in biological evidence
Black-box models
Outputs cannot be traced to biological evidence. Limited regulatory defensibility
Every output is traceable, evidence-linked, and auditable. Built for regulatory-grade scrutiny
The critical distinction: when independent analyses produce divergent results under identical conditions, they are sampling variability — not modeling biological constraint. PROTEUS V3 resolves this by measuring whether deterministic convergence occurs, distinguishing signal from noise at the architectural level.
DETERMINISTIC CONVERGENCE: THE CORE CONCEPT
Deterministic convergence is the reproducible emergence of stable multi-variant interaction configurations across independent simulations under identical conditions. It is not an assumption — it is a measured property of the biological system under investigation.
PROTEUS V3 does not impose convergence. It evaluates whether convergence occurs, and reports both outcomes explicitly:
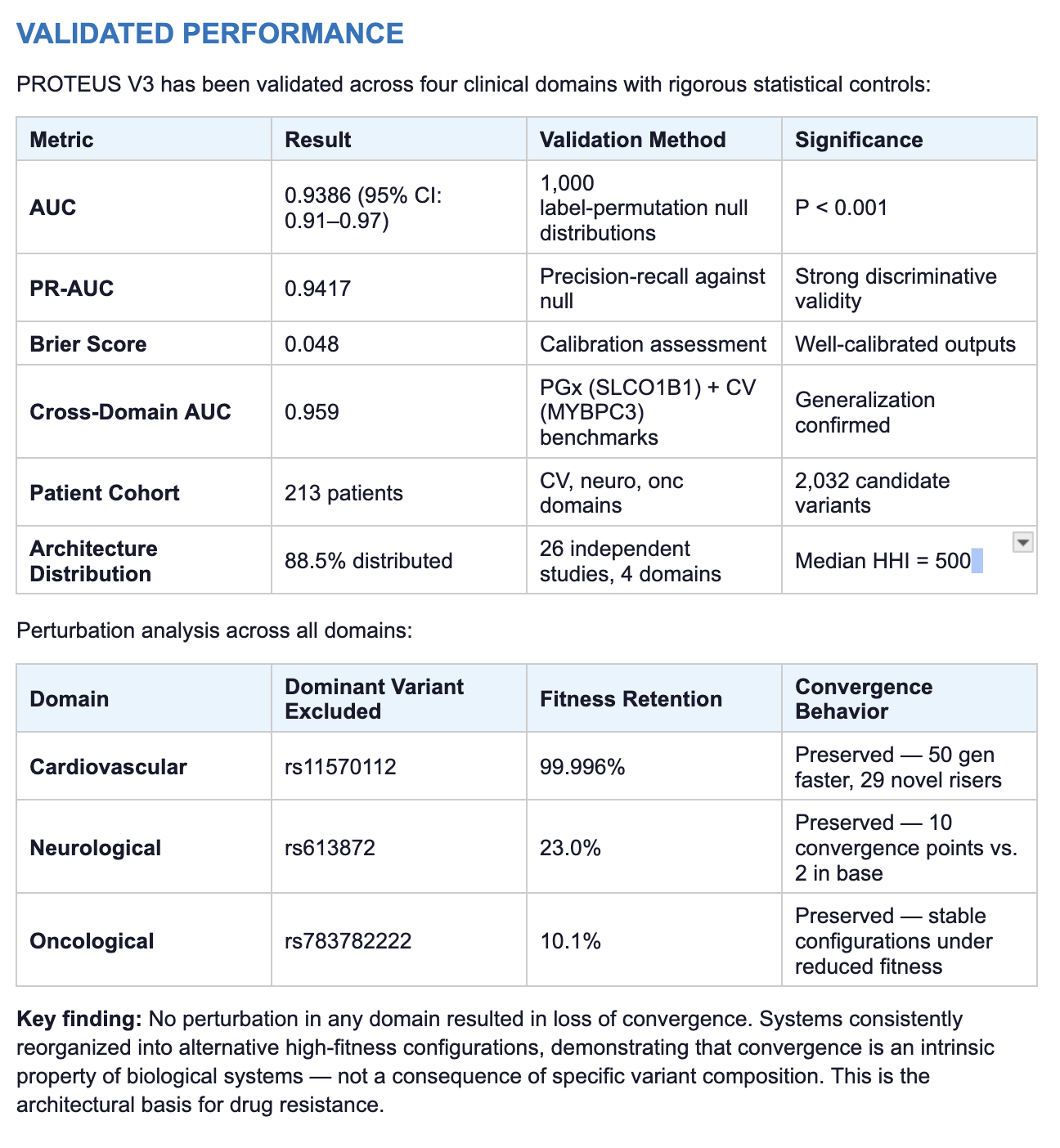
Reducible Systems
Distributed Systems
Converge toward a dominant interaction configuration with high stability across independent runs.
Characteristics:
95%+ recurrence of dominant interaction patterns
Low heterogeneity (I² ≈ 0)
Stable convergence within defined generation windows
High cross-run concordance of variant composition
Drug development implication: target selection is structurally grounded. Single-target strategies may be appropriate.
Do not collapse to a single solution. Multiple stable configurations persist under identical conditions.
Characteristics:
Multiple high-fitness configurations coexist
Rank-order varies across independent runs
System reorganizes around alternative pathways under perturbation
Removal of dominant variants does not cause collapse
Drug development implication: single-target strategies will face resistance through architectural reconfiguration. Combination approaches required.
This distinction is treated as a property of the biological system — not a failure of the model. Knowing whether a drug target sits within a reducible or distributed system fundamentally changes development strategy, resistance anticipation, and patient stratification.
WHERE PROTEUS V3 FITS IN THE DRUG DEVELOPMENT PIPELINE
1. Target Discovery & Validation
Identify genomic configurations associated with drug response variability through deterministic convergence analysis
Detect constrained interaction patterns across pathways that single-variant analysis misses
Distinguish reducible targets (structurally grounded) from distributed systems (requiring combination approaches)
Prioritize targets with stronger biological constraint evidence, not just statistical association
2. Preclinical Risk Assessment
Simulate potential resistance pathways under therapeutic pressure prior to in vitro testing
Identify high-risk genomic backgrounds where distributed architecture predicts treatment escape
Model reconfiguration trajectories — which alternative variant configurations emerge when dominant targets are suppressed
Reduce experimental trial-and-error by focusing preclinical resources on architecturally validated hypotheses
3. Clinical Trial Design & Patient Stratification
Stratify patient populations based on multi-variant interaction profiles, not individual biomarker status
Inform inclusion/exclusion criteria using constraint architecture classification (reducible vs. distributed)
Anticipate variability in drug response by identifying patients whose genomic architecture predicts reconfiguration under therapeutic pressure
Design adaptive trial protocols that account for distributed resistance mechanisms
4. Companion Diagnostic Development
Develop multi-variant CDx strategies that capture the distributed constraint architecture, not just dominant biomarkers
Identify why single-biomarker CDx misclassifies responders — concentration artifacts vs. structural vulnerability
Provide deterministic, reproducible evidence for CDx regulatory submissions
5. Post-Market Surveillance & Lifecycle Management
Model emerging resistance patterns in real-world patient populations
Support label expansion strategies by identifying additional patient populations with favorable constraint architectures
Inform next-generation drug design by mapping the distributed pathways tumors reconfigure through under first-generation therapeutic pressure

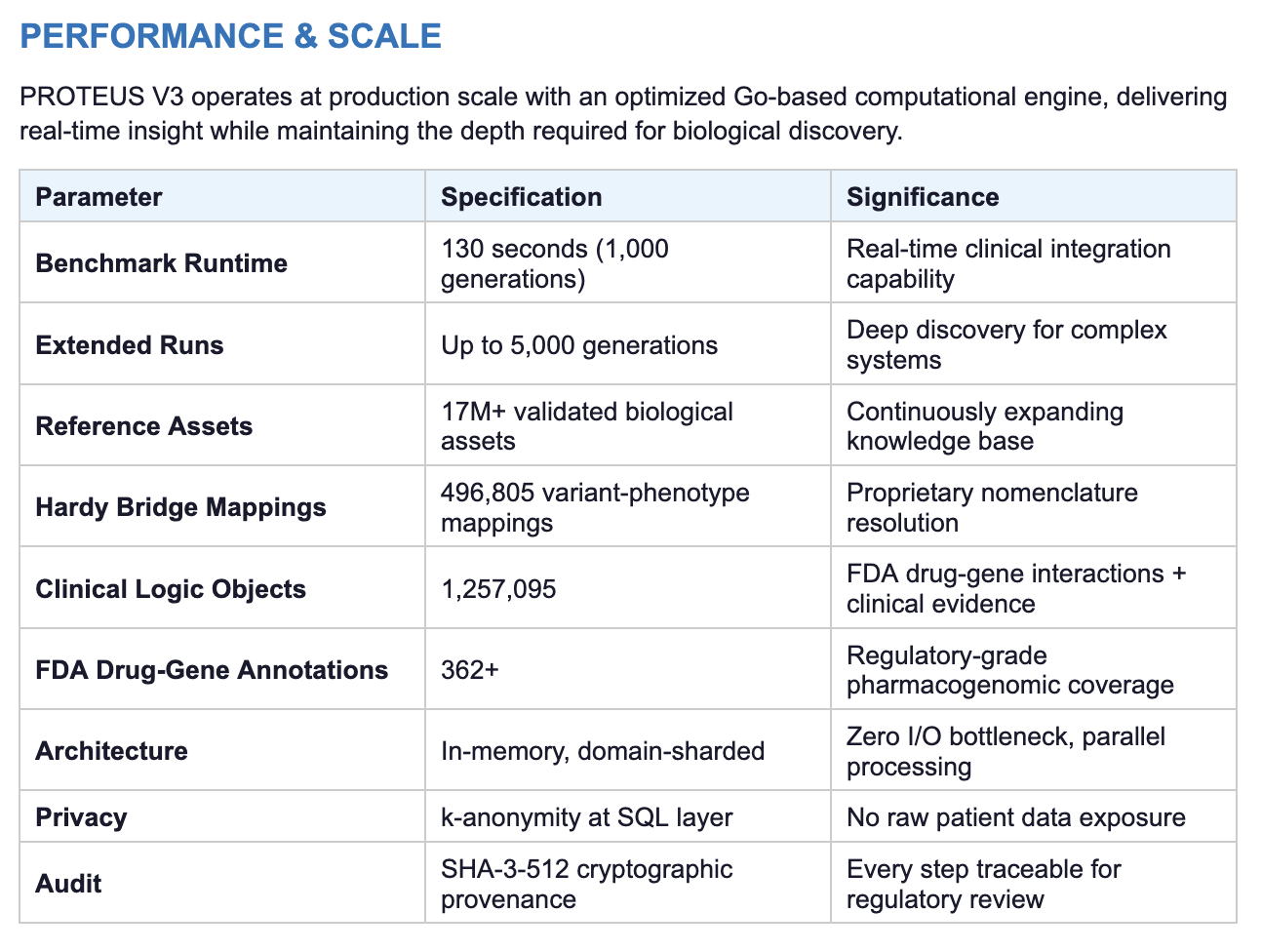
SCIENTIFIC FOUNDATION
Hardy, M.L. (2026). Deterministic Evolutionary Simulation Reveals Distributed Constraint Architectures in Human Genomic Systems. Submitted to Nature Biology. NomosLogic Inc., Salt Lake City, Utah.
NomosLogic holds 14 filed patent pillars protecting the Hardy Bridge nomenclature resolution system, the TRINITY multi-omic fusion engine, the PROTEUS evolutionary simulation framework, and the deterministic clinical logic architecture. Legal counsel: Wilson Sonsini Goodrich & Rosati. Delaware C-Corp.
ENGAGEMENT MODEL
PROTEUS V3 integrates into pharmaceutical workflows through flexible engagement structures:
Model
Description
Collaborative Research
Joint discovery programs applying PROTEUS V3 to specific therapeutic areas or pipeline targets. Defined scope, shared milestones, IP terms structured per engagement.
Targeted Discovery Studies
Focused analyses of specific drug targets, resistance mechanisms, or patient stratification challenges. PROTEUS V3 processes your genomic data and delivers deterministic convergence analysis, constraint architecture classification, and perturbation modeling.
Platform Licensing
License PROTEUS V3, Hardy Bridge, and TRINITY engines for integration into internal discovery and development pipelines. Includes access to 17M+ reference assets and ongoing platform updates.
CDx Co-Development
Joint development of interaction-aware companion diagnostics that capture distributed constraint architectures. Deterministic outputs designed for regulatory submission.
THE FUNDAMENTAL QUESTION
Drug discovery does not fail due to lack of data. It fails when analytical systems cannot model biological complexity, reproduce results, or distinguish constraint from variability.
PROTEUS V3 addresses these failures directly. By measuring deterministic convergence, it classifies genomic systems as constrained or distributed — shifting drug discovery from association-driven inference to reproducible, structure-aware biological discovery.
The question is not whether genomic architecture is distributed — our data demonstrate that it is, across 88.5% of observed systems. The question is whether your pipeline has the infrastructure to leverage that reality before it becomes a late-stage failure.
Start the Conversation
Matt Hardy, CEO · NomosLogic, Inc.