Deterministic Convergence: Why Genomic Systems Don’t Break When You Remove Their Most Important Parts
For decades, genetics has largely been framed around a simple idea:
find the important gene, understand the trait.
That model has been productive—but incomplete.
What we are now seeing suggests something deeper.
A different behavior emerges
Across multiple independent domains—cardiovascular, neurological, oncological, and now renal—we observe a consistent pattern:
When the most influential genetic variants are removed, the system does not collapse.
Instead, it reorganizes.
New variants rise.
Alternative pathways activate.
The system finds another way to reach a stable state.
And most importantly:
it still converges.
What we’re calling deterministic convergence
We use the term deterministic convergence to describe this phenomenon:
Under identical conditions, complex genomic systems repeatedly evolve toward stable, reproducible configurations—even when key components are removed.
This is not random recovery.
This is not statistical noise.
This is structured behavior.
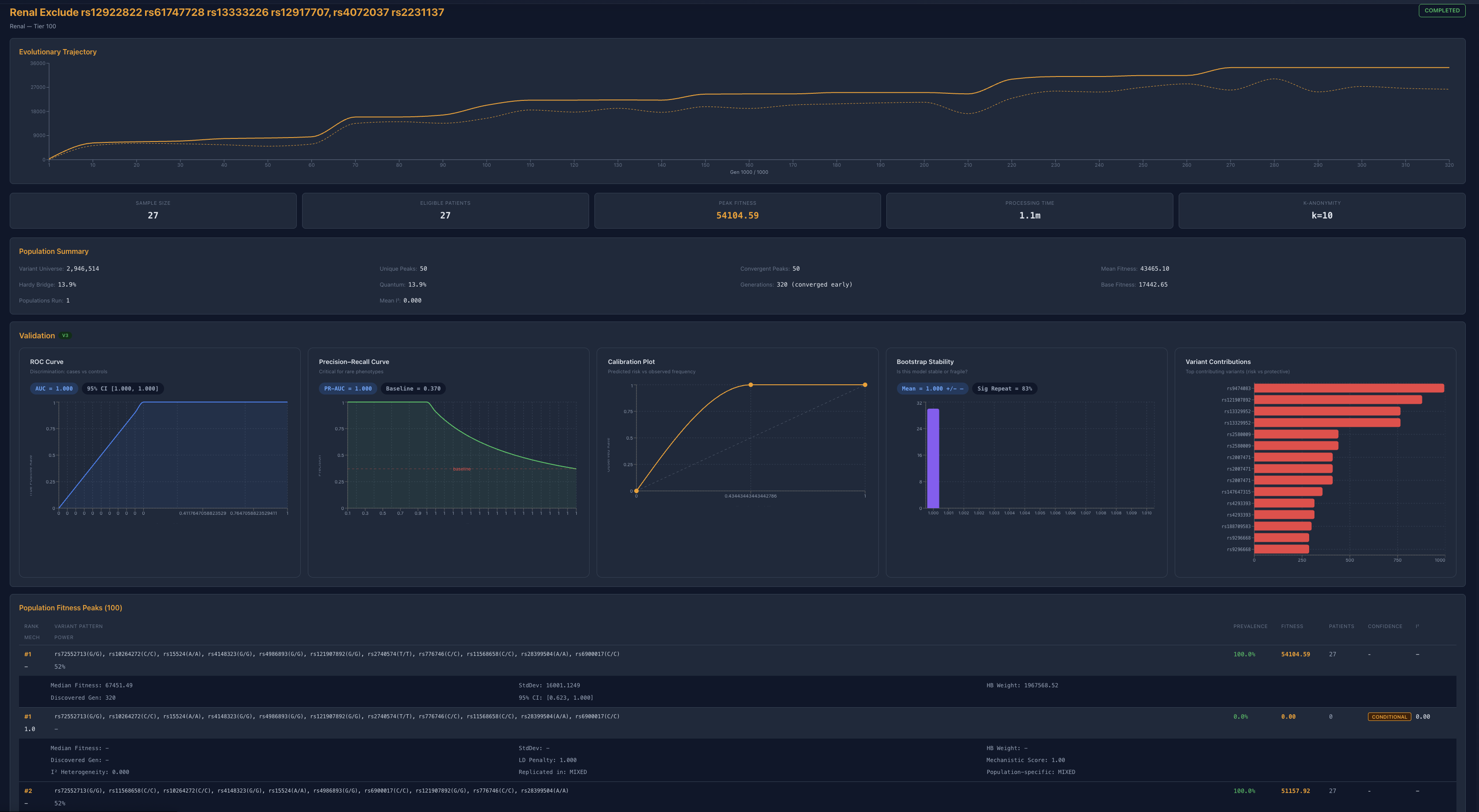
The renal system makes this impossible to ignore
In the renal domain, we removed variants strongly associated with kidney function:
UMOD-related variants (linked to eGFR and hypertension)
NPHS2 (podocin) variants (linked to nephrotic syndromes)
Additional top-ranked contributors
These are not trivial signals. These are biologically meaningful anchors.
And yet:
The system retained high fitness
Convergence still occurred
Entirely new variant combinations emerged
In some cases, convergence even happened faster after removing dominant variants.
That shouldn’t happen under a linear model
If traits were driven primarily by a small number of dominant variants, removing them should degrade performance or prevent convergence.
It doesn’t.
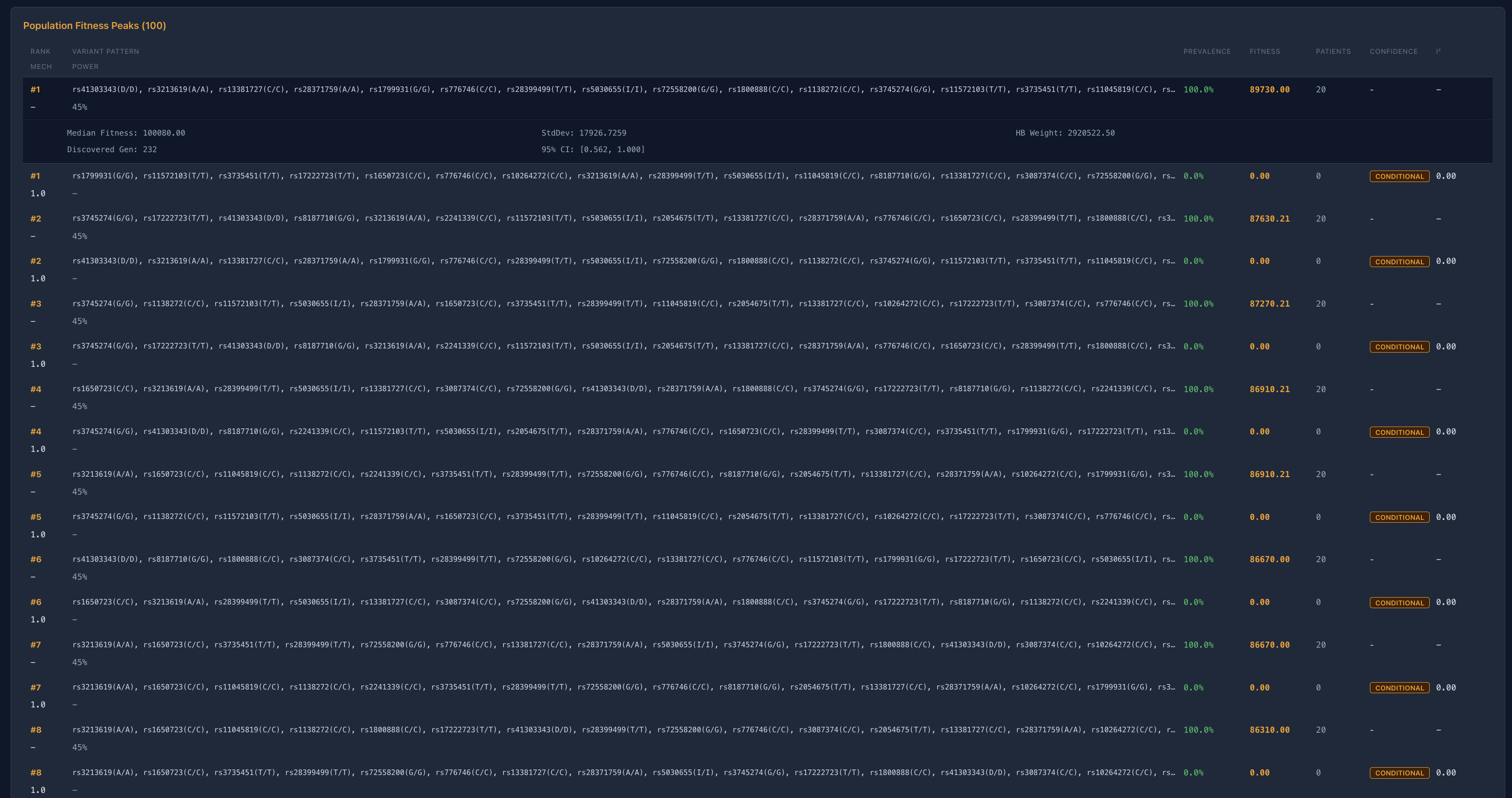
Instead, what we observe is:
multiple interchangeable high-fitness configurations
That is not a single-path system.
That is a distributed system.
From “genes cause traits” to “systems resolve constraints”
What these results suggest is a shift in perspective:
Instead of:
genes → traits
We may need to think in terms of:
systems → constraint satisfaction → stable configurations
Traits are not the output of a single causal chain.
They are the result of a system navigating a landscape of possibilities—and settling into one of several viable solutions.
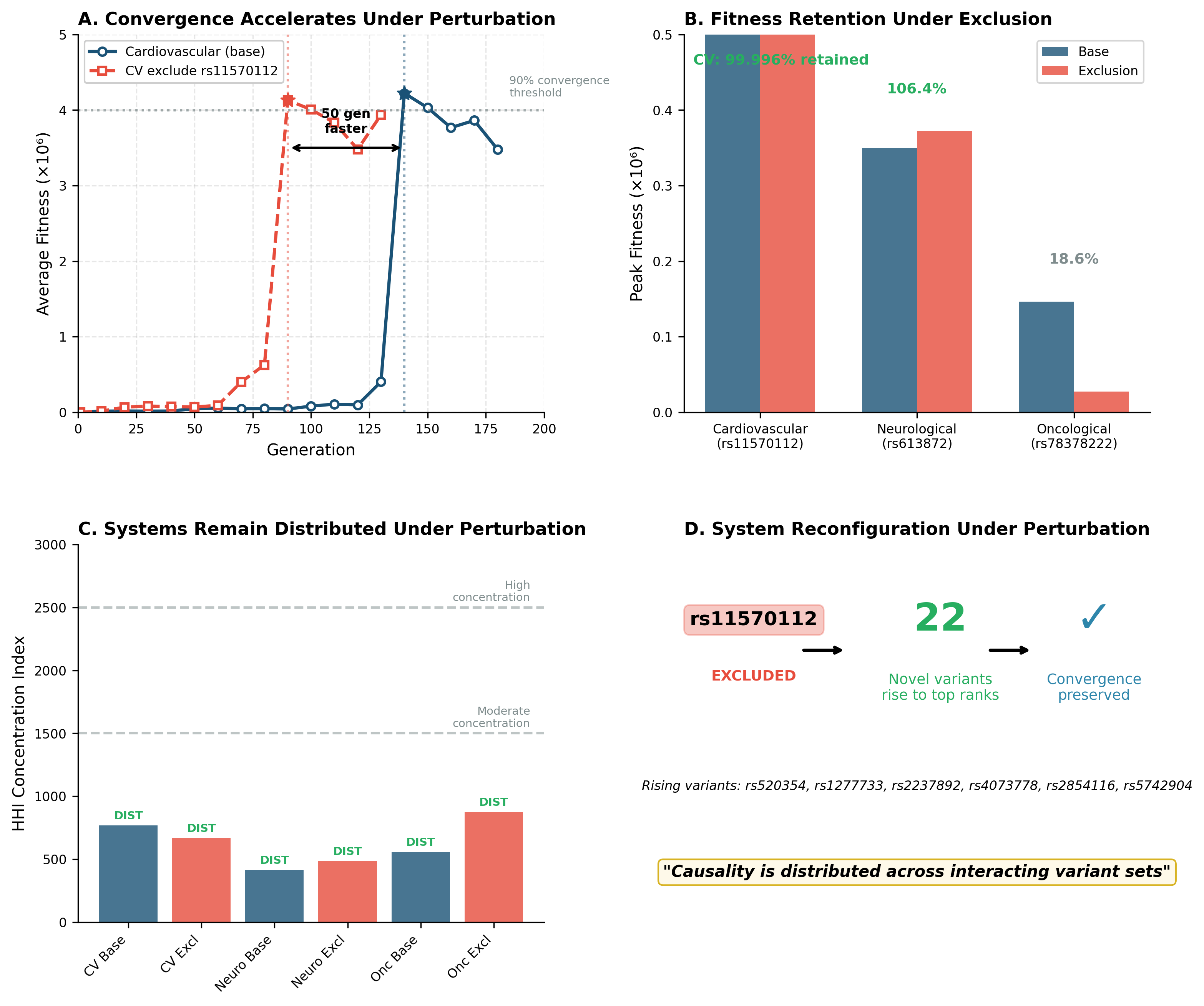
Why removing variants sometimes helps
One of the most surprising findings is that removing dominant variants can sometimes accelerate convergence.
This suggests something subtle but important:
Dominant variants may act as anchors, concentrating the system
Removing them may free the system to explore alternative configurations
The system then converges through a different, sometimes more efficient pathway
This is not failure.
This is adaptability.
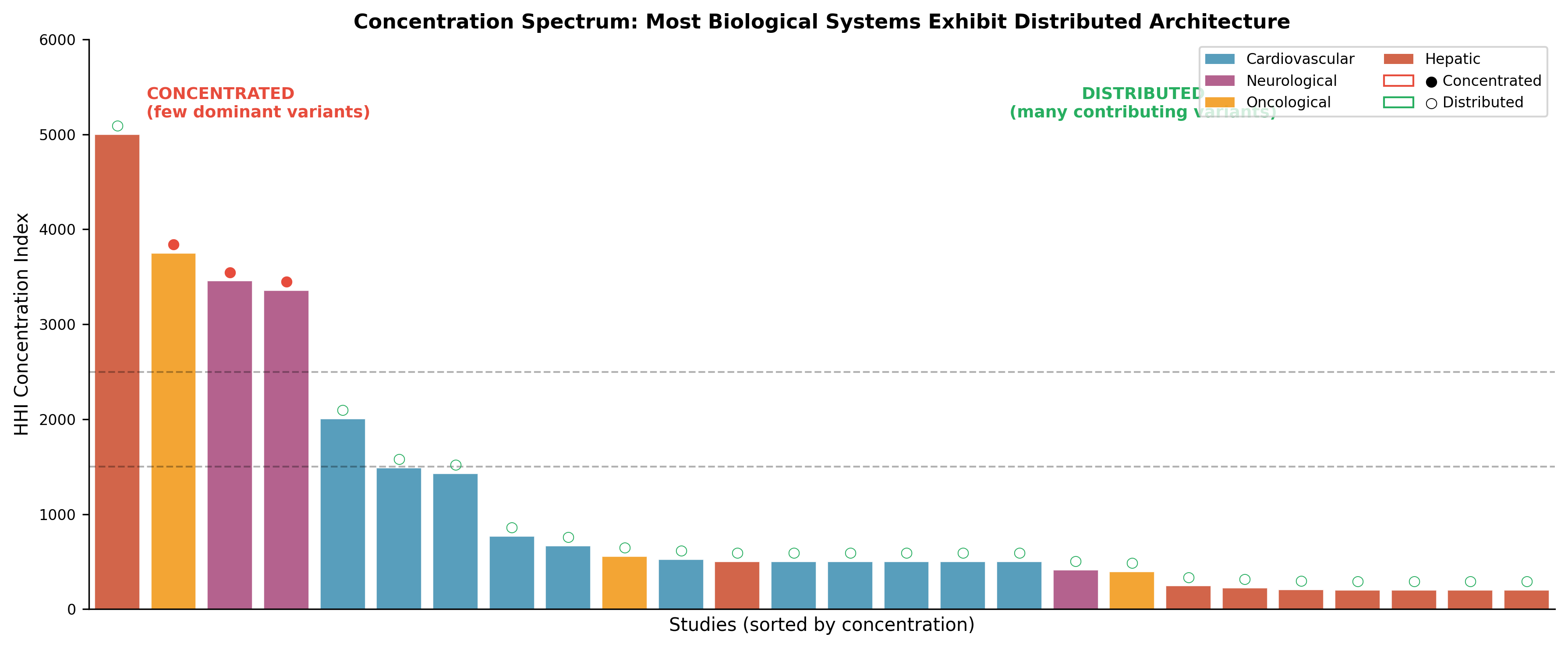
A distributed architecture
Across all domains studied so far, we see the same structural property:
Causality is not concentrated—it is distributed across interacting variant sets.
No single variant is strictly required.
What matters is the configuration, not the individual component.
What this means going forward
If this holds up under continued validation, it has implications across multiple areas:
Genomic research: shifts focus from single-variant effects to system-level interactions
Drug discovery: targeting pathways instead of individual genes becomes more natural
Precision medicine: multiple equivalent intervention strategies may exist for the same phenotype
But more fundamentally:
it changes how we think about biological systems themselves
Where this goes next
This is not a finished story.
It’s the beginning of one.
The goal now is not to declare a new principle prematurely—but to test it, stress it, and attempt to break it.
So far, across multiple domains:
it hasn’t broken.
A simple way to explain it
If you had to explain this to someone outside of science, it would be this:
Biology doesn’t rely on a single solution—it finds one of many.
And even when you remove what looks like the most important piece:
it finds another way.
Final thought
We may have been asking the wrong question.
Not:
“Which gene causes this trait?”
But:
“What configurations can this system settle into?”
Deterministic convergence suggests:
there are more answers than we thought.
Appendix:
— Matthew Hardy
NomosLogic