Abstract
Current paradigm: Polygenic risk assumes additive, independent variant effects.
Proteus V3 paradigm: Biological systems are organized into interaction hierarchies—some collapse to single dominant drivers, while others depend on coordinated network integrity.
We applied an evolutionary optimization framework (Proteus V3) across multiple biological domains—cardiovascular, oncological, metabolic, and neurological—to identify stable, multi-variant genetic interaction structures. Across all domains, the system consistently converged on low-dimensional, interpretable hierarchies composed of dominant variants (“anchors”), secondary modifiers, and non-contributing conditional variants.
Two distinct modes of genomic organization emerged:
Reducible systems, characterized by a single dominant genetic axis (e.g., MYBPC3 in cardiovascular, TP53 in oncology)
Distributed systems, requiring multiple co-dependent variants to explain observed structure (e.g., PRNP–TCF4–lysosomal axis in neurology)
These findings suggest that complex genomic phenotypes may be representable as hierarchical interaction structures, and that biological systems differ in their degree of reducibility. This framework provides a foundation for interpretable, system-level genomic analysis.
Introduction
Genomic interpretation has traditionally relied on:
single-variant associations (GWAS)
additive models (polygenic risk scores)
isolated pharmacogenomic rules
These approaches fail to capture:
conditional dependencies between variants
hierarchical importance
system-level organization
To address this, we developed Proteus V3, an evolutionary optimization system that:
searches for combinatorial variant sets
evaluates them using a composite fitness function
validates findings via cross-validation, permutation testing, and bootstrap stability
This study investigates whether consistent structural patterns emerge across biological domains, and whether genomic systems exhibit differing degrees of dimensional reducibility.
Methods
Cohorts
Multiple domain-specific cohorts were analyzed:
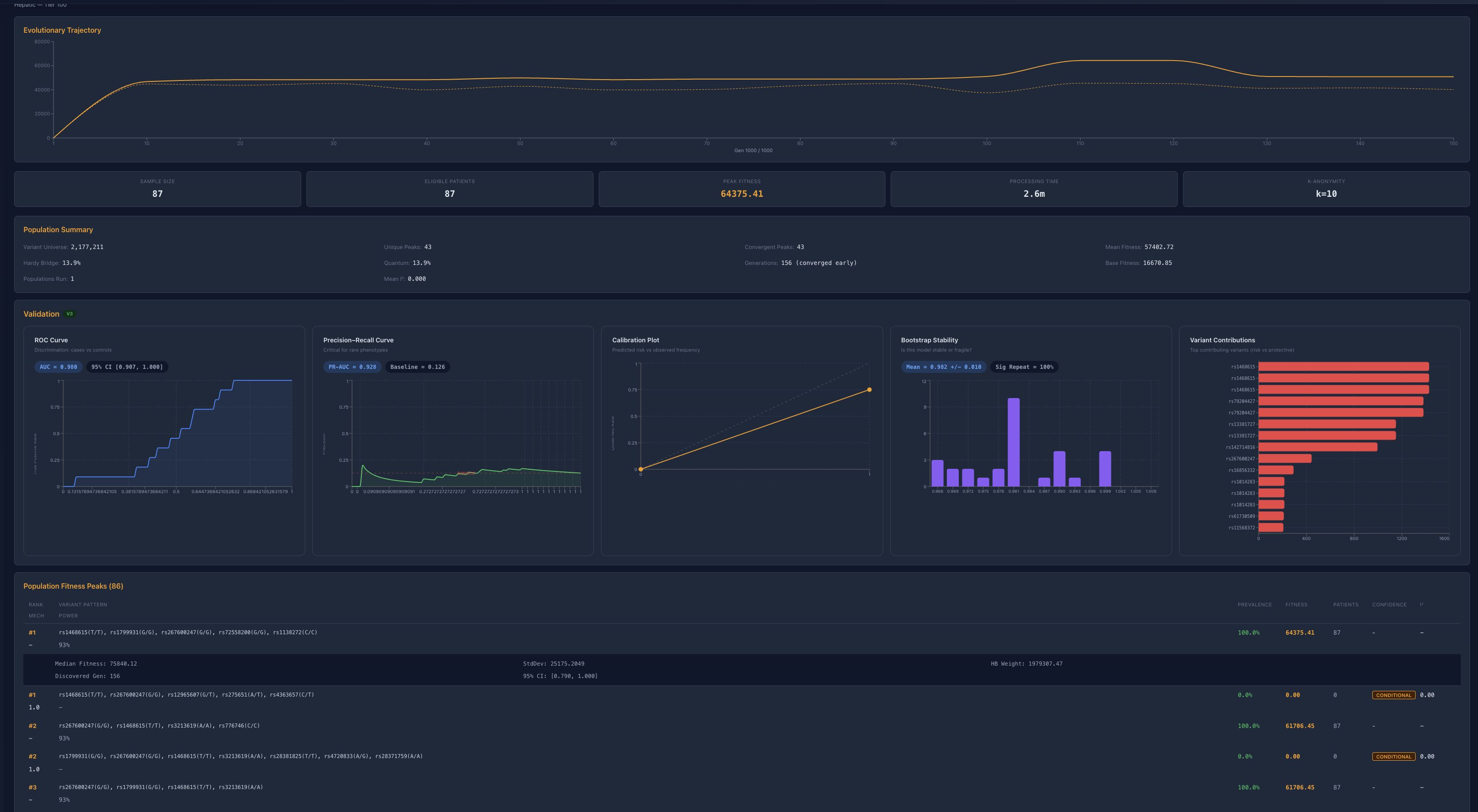
Cardiovascular (n≈87)
Oncological (n≈13)
Neurological (n≈12)
Metabolic (similar scale)
Each cohort included:
genotype data (~1.5M–2.1M variants)
phenotype labels
clinical weight mappings
Evolutionary Optimization
Proteus V3 uses a genetic algorithm to evolve variant combinations (“fitness peaks”) that maximize:
clinical effect size
mechanistic pathway relevance (KEGG, Reactome, PPI)
linkage disequilibrium penalties
prevalence and stability
Validation Framework
Each run includes:
k-fold cross-validation
permutation testing (null distribution)
bootstrap resampling (stability)
Metrics:
AUC / PR-AUC
calibration
peak recurrence frequency
fitness variance
Results
1. Consistent Emergence of Interaction Hierarchies
Across all domains, Proteus V3 converged on a shared structural pattern:
Three-layer hierarchy
Anchor layer – dominant variant(s)
Modifier layer – secondary contributors
Non-contributing layer – rejected conditional variants
This structure was:
stable across runs
reproducible under resampling
resistant to overfitting (validated via permutation)
2. Reducible Systems (Single-Anchor Behavior)
Cardiovascular Domain
Anchor: MYBPC3
Modifier: CYP2C19
Additional variants → minimal incremental fitness
System collapses to a single dominant structural axis
Oncological Domain
Anchor: TP53 (rs78378222)
Secondary variants → minor contributions
Strong convergence to a master regulatory gene
Key property
These systems exhibit:
Low-dimensional reducibility
→ phenotype largely explained by a single dominant variable
3. Distributed Systems (Multi-Anchor Behavior)
Neurological Domain
Core cluster:
PRNP (protein folding)
TCF4 (transcriptional regulation)
Lysosomal/degradation-associated variants
Characteristics:
multiple variants required for maximal fitness
no single variant dominates completely
system does not collapse under simplification
Key property
Distributed dependency structure
→ phenotype requires multiple co-equal biological axes
4. Conditional Variant Rejection
Across all domains:
Over-specified variant combinations → 0% prevalence
No reproducibility
Zero fitness contribution
👉 The system consistently prunes:
biologically implausible combinations
statistical artifacts
5. Stability and Convergence
Across domains:
rapid convergence (<250 generations)
low fitness variance
high bootstrap repeat rates (>80–100%)
👉 Indicates:
strong signal
low stochastic noise
stable solution landscapes
Discussion
1. Emergence of Genomic Reducibility
This study introduces the concept of:
Genomic reducibility
Defined as:
the degree to which a system can be explained by a small number of variants
Two observed regimes:
Type. Description. Example
Reducible. Single dominant axis. MYBPC3, TP53
Distributed. Multi-axis dependency. PRNP + TCF4 cluster
2. Biological Interpretation
Reducible systems:
dominated by:
structural constraints (cardio)
master regulators (oncology)
Distributed systems:
require:
simultaneous integrity across pathways
network-level stability (neurology)
3. System-Level Insight
Proteus V3 does not merely identify variants.
It identifies:
how biological systems organize their genetic dependencies
4. Implications
Scientific
Enables identification of interaction hierarchies
Moves beyond additive genetic models
Provides testable hypotheses about system organization
Clinical (future)
Patient stratification based on interaction structures
Identification of dominant vs conditional drivers
Hypothesis generation for therapeutic targeting
Conclusion
Proteus V3 consistently identifies low-dimensional genomic interaction hierarchies across biological domains. These hierarchies reveal whether systems are:
reducible (single dominant axis), or
distributed (multi-axis dependency)
This suggests that complex genomic phenotypes may be representable as structured, hierarchical systems, providing a new framework for interpretable genomic analysis.