Editorial
For decades, the dominant paradigm in genomics has been implicitly simple: identify the variants most strongly associated with a trait, and you are closer to identifying its cause.
But this assumption carries a hidden expectation — that biological systems are, at some level, reducible. That removing the most influential component should degrade or collapse the system’s behavior.
What if that assumption is wrong?
The problem: instability disguised as insight
Many current approaches to complex trait analysis rely on probabilistic or statistical association. While powerful, these methods often exhibit instability:
Results vary across cohorts
Models shift with new data
Repeated analyses do not always yield consistent structures
If the underlying system is stable, why are our models not?
The observation: systems converge
In controlled simulation studies of genomic interaction networks, a different pattern emerges.
Across independent runs under identical conditions, systems repeatedly converge toward stable configurations of interacting variants. These configurations are not identical in composition, but they are consistent in structure and behavior.
This phenomenon can be described as deterministic convergence:
The reproducible emergence of stable interaction structures across independent simulations under identical conditions.
The critical test: perturbation
A key question follows:
If these systems are truly driven by dominant variants, what happens when those variants are removed?
The expectation under a traditional model is straightforward: disruption, degradation, or collapse.
But that is not what is observed.
Instead:
Systems continue to converge
Fitness landscapes reorganize
Previously lower-ranked variants rise to prominence
Overall system behavior is preserved
In some cases, convergence occurs more rapidly after perturbation.
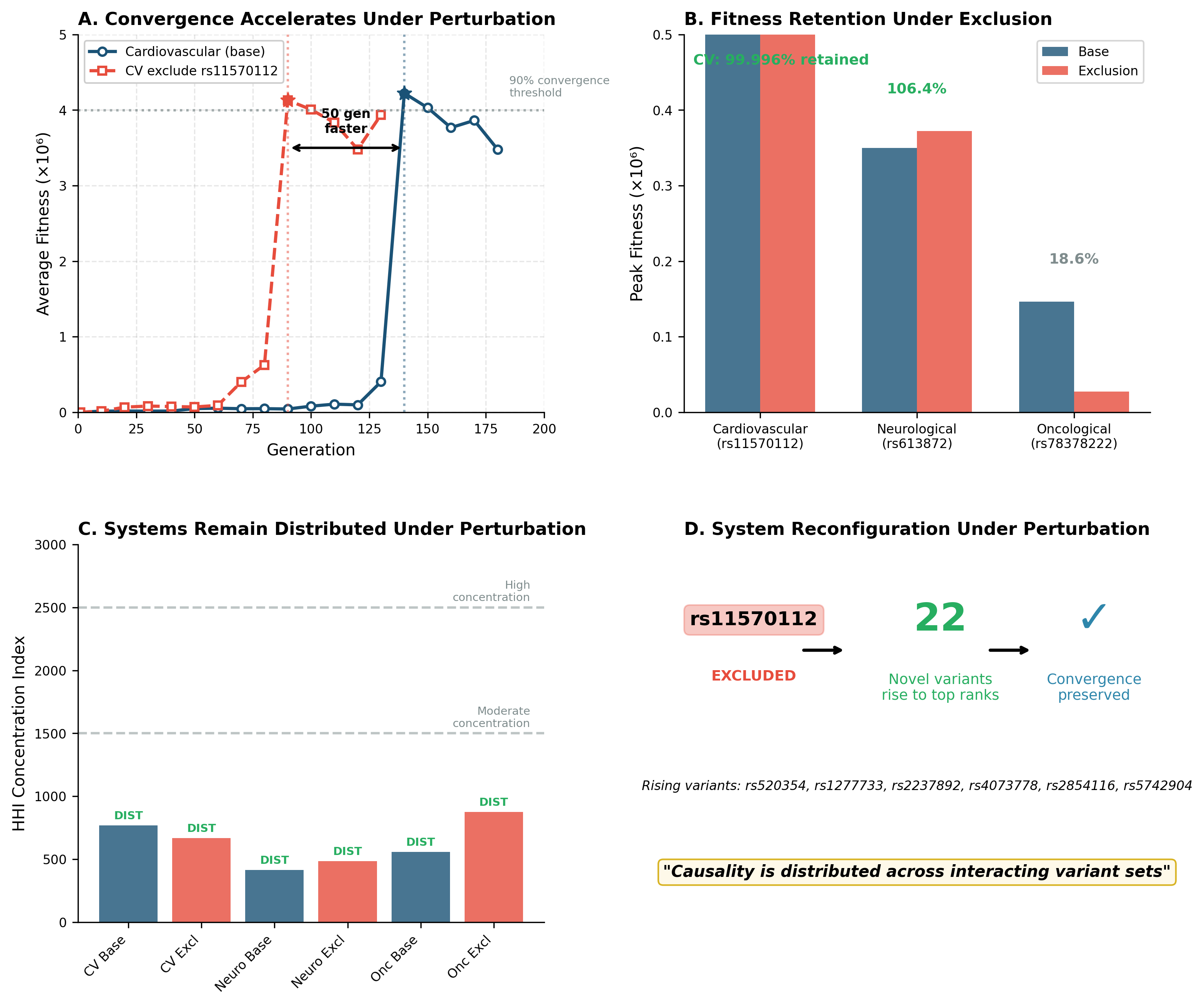
The implication: causality is distributed
These observations suggest a different model of biological organization.
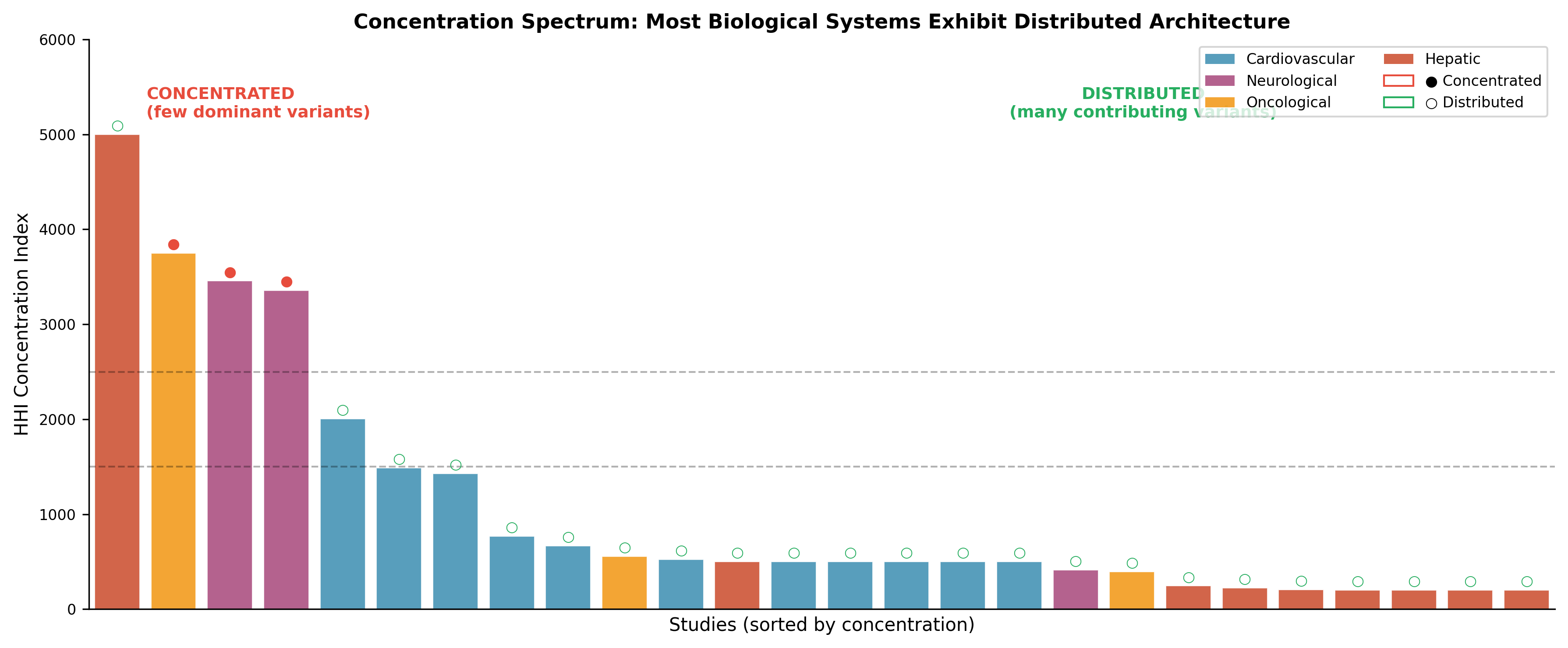
Rather than being governed by a small number of dominant causal variants, complex traits appear to be structured as distributed constraint systems:
Multiple variants contribute collectively
Stability arises from interaction patterns, not individual components
Apparent “dominance” reflects concentration, not necessity
When a highly influential variant is removed, the system does not fail — it reconfigures along alternative constraint pathways.
A shift in perspective
This has important implications.
If biological systems are distributed:
The question is not “which variant causes disease?”
But rather “how do interacting variants constrain the system toward stable outcomes?”
This reframing moves away from reductionist causality and toward system-level behavior.
Limits and scope
Not all biological systems are expected to exhibit deterministic convergence.
Some may remain:
highly context-dependent
multi-stable
or non-convergent under certain conditions
Distinguishing between convergent and non-convergent architectures becomes an important problem in itself.
Where this leads
If convergence is a real and generalizable property of biological systems, it suggests that we are not simply modeling biology — we are beginning to observe its underlying structure.
And that structure may be less about isolated causes, and more about constraints that shape possible outcomes.



