Executive Summary
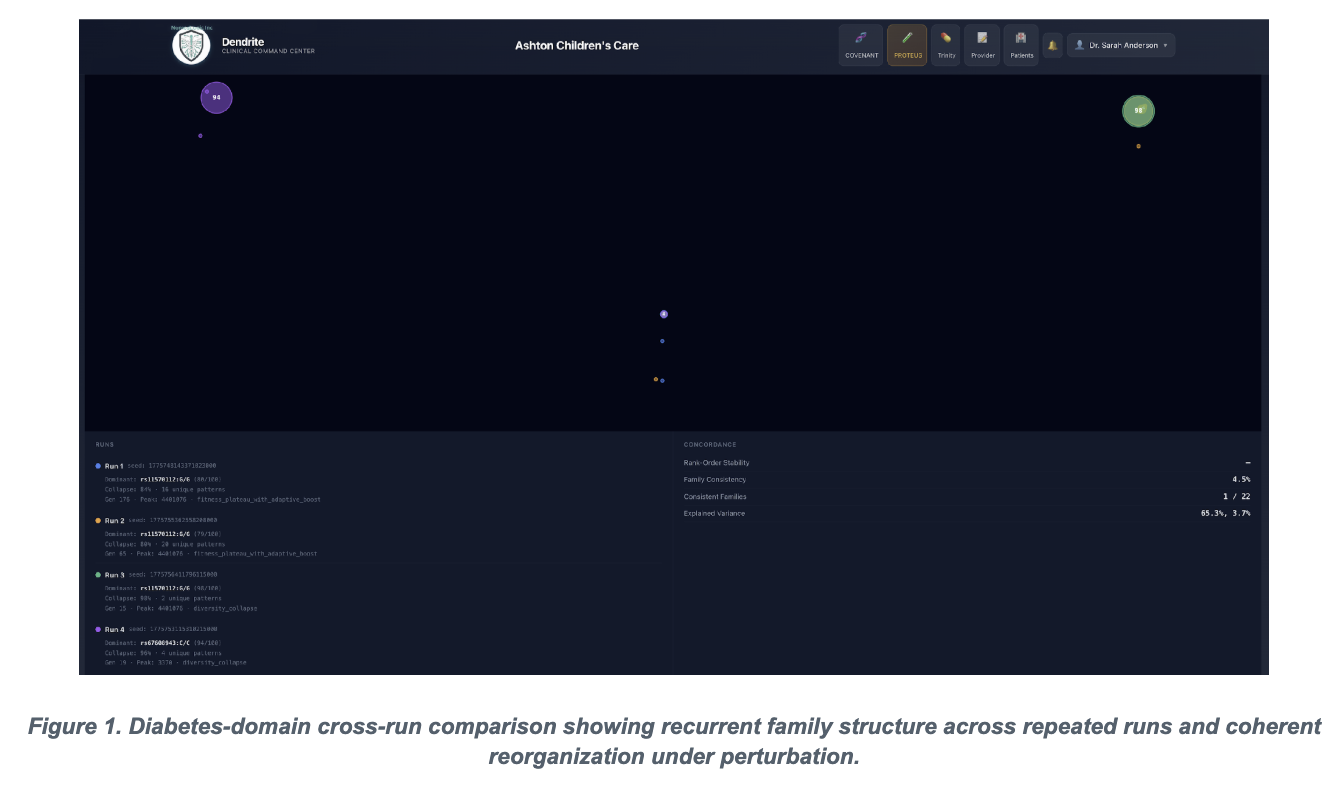
The submitted PROTEUS manuscript argues that complex genomic systems do not behave like fragile single-locus architectures. Across six clinical domains, the study reports reproducible convergence under identical conditions, structured reconfiguration under targeted perturbation, and a finite collapse boundary in hematology only after progressive multi-locus exclusion. In practical terms, the paper reframes genomic risk architecture as distributed, layered, and compensatory rather than dominated by isolated points of failure.
Why This Paper Matters
The paper is not simply introducing a new simulation engine. It is using repeated replication and targeted exclusion studies to test a more fundamental question: whether phenotype-associated genomic systems resolve toward stable interaction families, and whether those families survive perturbation by reorganizing instead of collapsing. That distinction matters because most translational genomics still treats associated variants as if they were individually decisive. If the underlying structure is actually distributed, single-target reasoning will systematically overestimate fragility and underestimate compensation.
What the Manuscript Shows
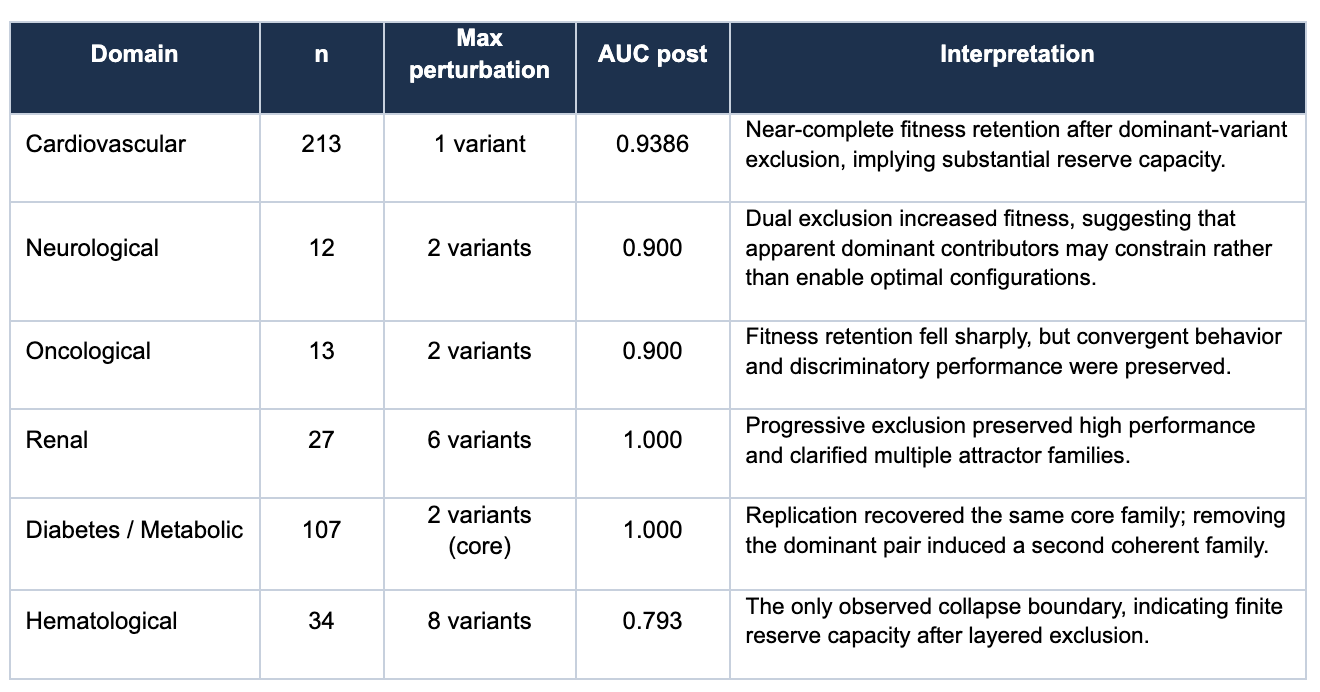
The manuscript presents six clinical domains spanning cardiovascular, neurological, oncological, renal, diabetes and metabolic, and hematological cohorts. Across 406 patients, the reported pattern is consistent: dominant contributors can be removed, fitness often remains high, and discriminative performance is preserved in five of six domains. The diabetes domain provides the clearest exact-parameter replication example, where repeated runs recover the same core family under fixed conditions. The hematological domain provides the clearest boundary example, where progressive exclusion eventually degrades the system into a lower-fidelity regime rather than a merely cosmetic reconfiguration.
How to Read the Central Claims
The strongest interpretation is not that the manuscript proves an absolute biological law. The strongest interpretation is that, under deterministic execution on real patient genomic data, the system exhibits reproducible attractor behavior and non-trivial reserve capacity across multiple domains. That is enough to challenge naive single-locus framing. It is also enough to support the more bounded claim that at least some genomic disease architectures are layered, compensatory, and structurally distributed.
Key Findings by Domain
Selected Figures
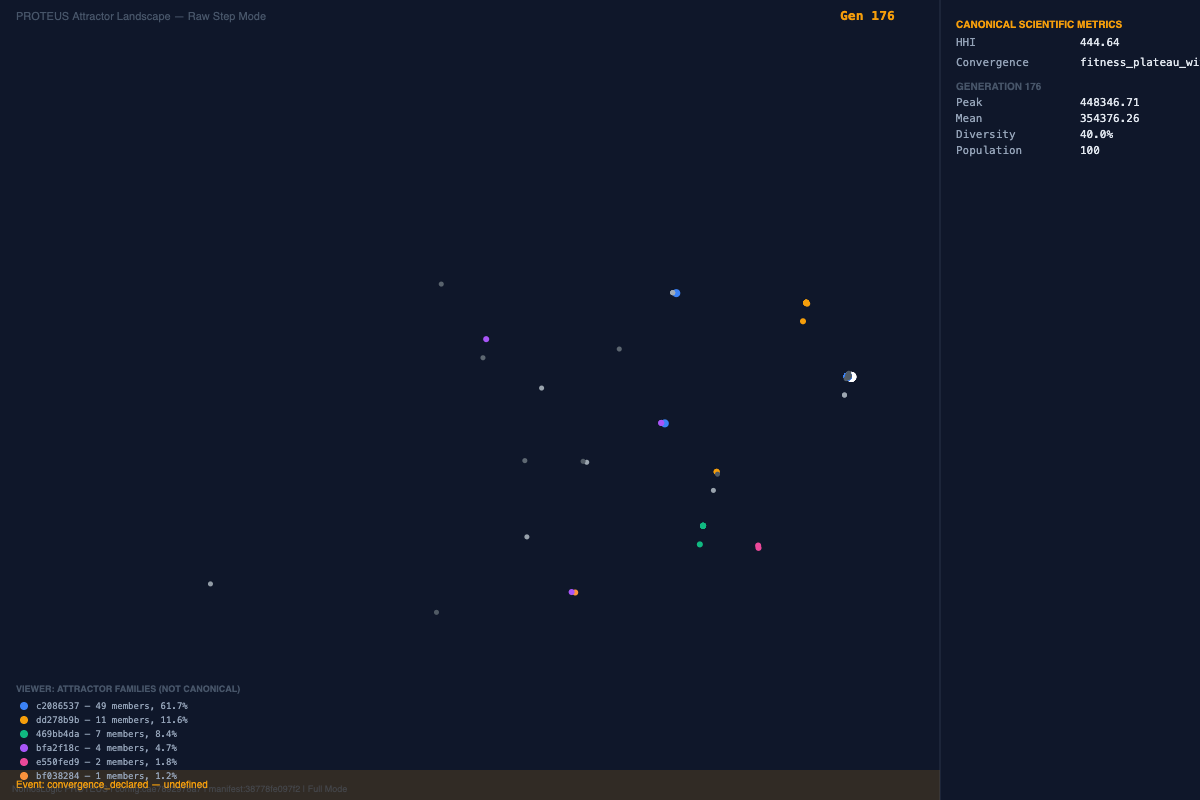
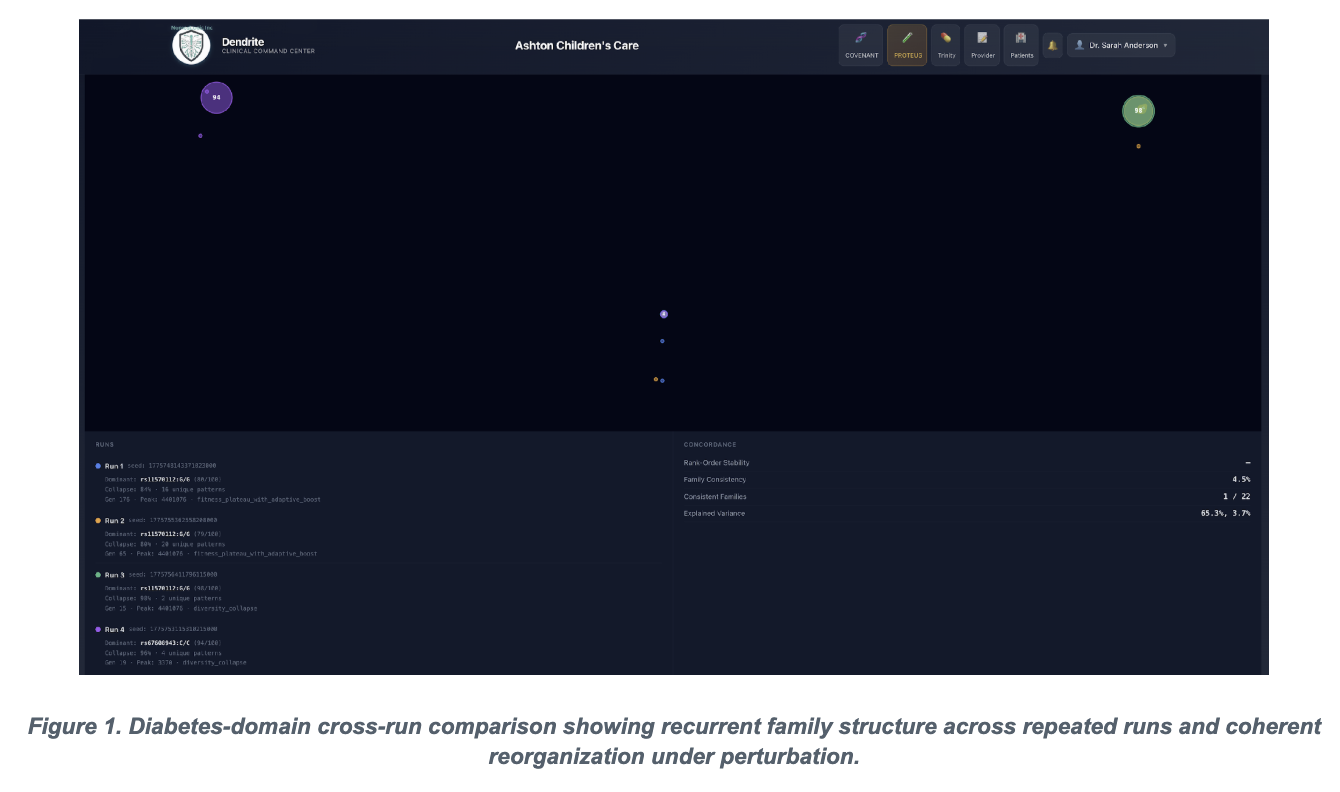
The figures below are included because they capture the manuscript's two strongest visual arguments: reproducible family-level reappearance in diabetes and finite reserve-capacity breakdown in hematology.
What Makes the Diabetes Domain Important
The diabetes and metabolic cohort functions as the paper's clearest proof-of-principle for deterministic convergence. The submitted manuscript reports that repeated runs under identical parameters recover the same core family around rs147834269 and rs1801212, with identical peak fitness in the replicate runs. That is the kind of result that separates a stochastic storytelling engine from a system that is actually discovering stable structures under controlled conditions.
Why the Hematology Series Matters
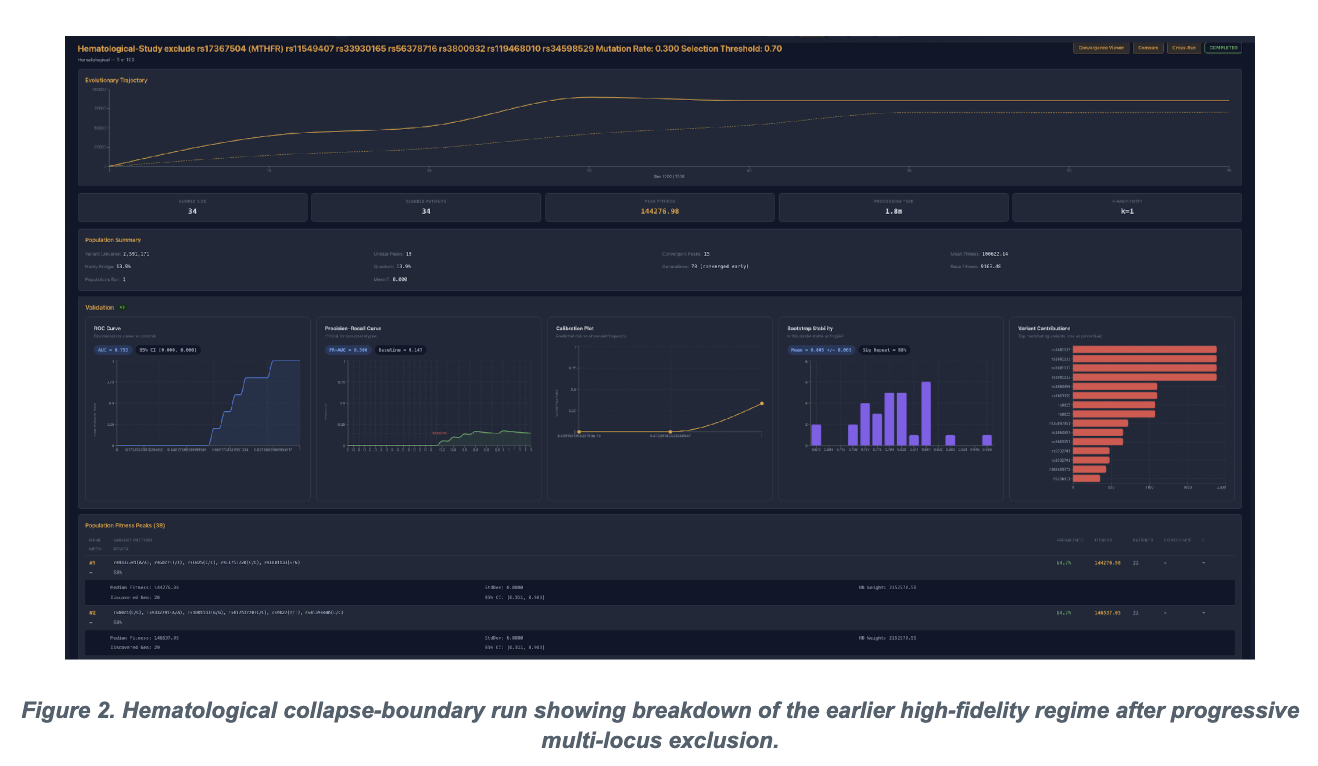
The hematology series is important because it prevents the paper from sounding rhetorical. The manuscript does not stop at saying systems reorganize. It pushes the architecture through sequential exclusion until the reserve capacity thins and a measurable collapse boundary appears. The inflection point is not just lower peak fitness. It is the simultaneous degradation in AUC, bootstrap stability, and the quality of the recovered basin. That is what gives the paper a finite boundary instead of a vague resilience narrative.
Implications for Drug Discovery and Genomic Interpretation
If genomic systems are distributed rather than concentrated, then intervention strategies aimed at apparently central nodes may face compensation rather than durable control. That implication touches therapeutic target selection, polygenic modeling, and cross-population replication. The practical message is not that single targets never matter. It is that target centrality should be tested against reserve architecture, not assumed from surface association alone.
What the Paper Does Not Claim
• The paper does not claim universal biological determinism.
• It does not claim that simulation output is equivalent to direct mechanistic proof.
• It does not claim that all domains share the same collapse boundary or reserve depth.
• It does claim that reproducible, perturbation-persistent system behavior is observable across multiple independent domains under deterministic execution.
Closing Perspective
This manuscript matters because it begins to move the conversation from association toward architecture. Its strongest contribution is not a slogan. It is a bounded empirical pattern: replicated convergence, persistent reconfiguration, and one observed collapse boundary after layered perturbation. That combination is enough to justify serious scientific attention, and it provides a clearer foundation for the next stage of validation, criticism, and industrial application.