For years, genomics has been treated primarily as an interpretation problem.
More data.
More models.
More prediction layers.
More attempts to extract signal from growing biological complexity.
But the central problem may be simpler than that.
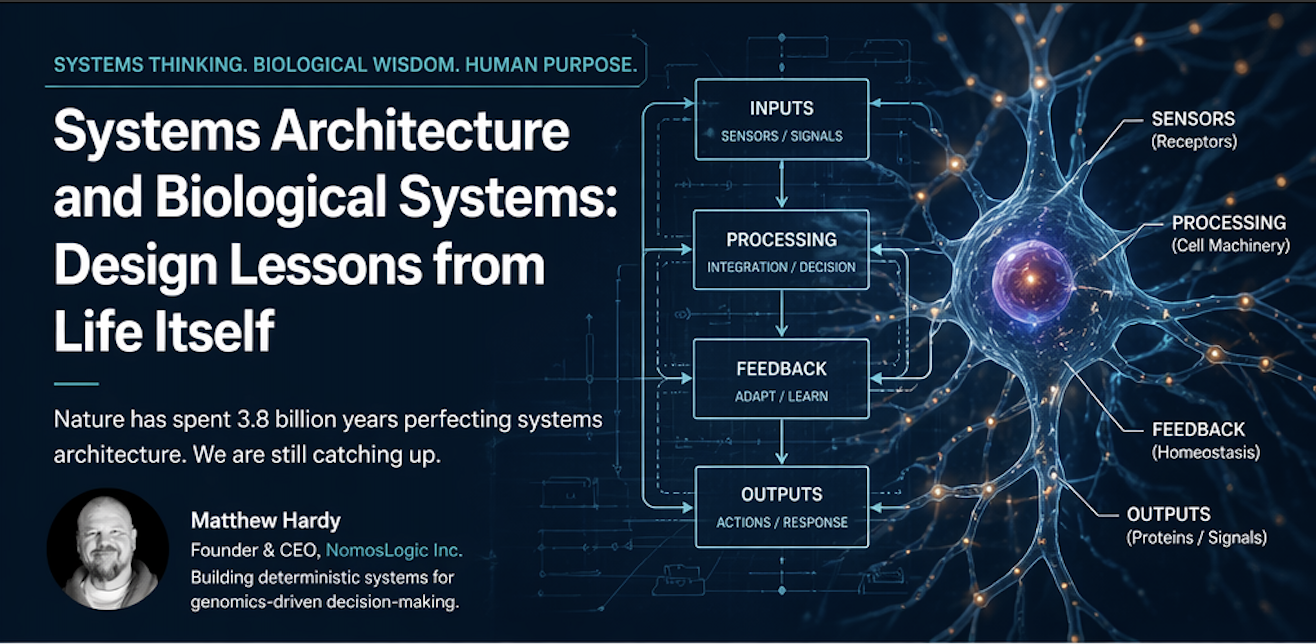
Genomics does not just have an analysis problem. It has a systems problem.
Modern genomic pipelines remain fragmented across incompatible data formats, inconsistent nomenclature, variable evidence layers, and inference systems that are often difficult to reproduce, inspect, or defend. In high-consequence settings, that matters. If the same biological input does not reliably resolve into the same governed output, the problem is not only scientific. It is architectural.
This is where systems engineering needs to enter the conversation.
In mature engineering domains, reliability does not come from adding more interpretive layers on top of unstable foundations. It comes from building infrastructure that is deterministic, traceable, and resilient under stress. Genomics should be no different.
That means moving beyond the idea that biological meaning lives only in isolated markers or static associations. Many of the most important behaviors in complex systems are not properties of single parts viewed alone. They emerge from interaction structure.
This has important consequences.
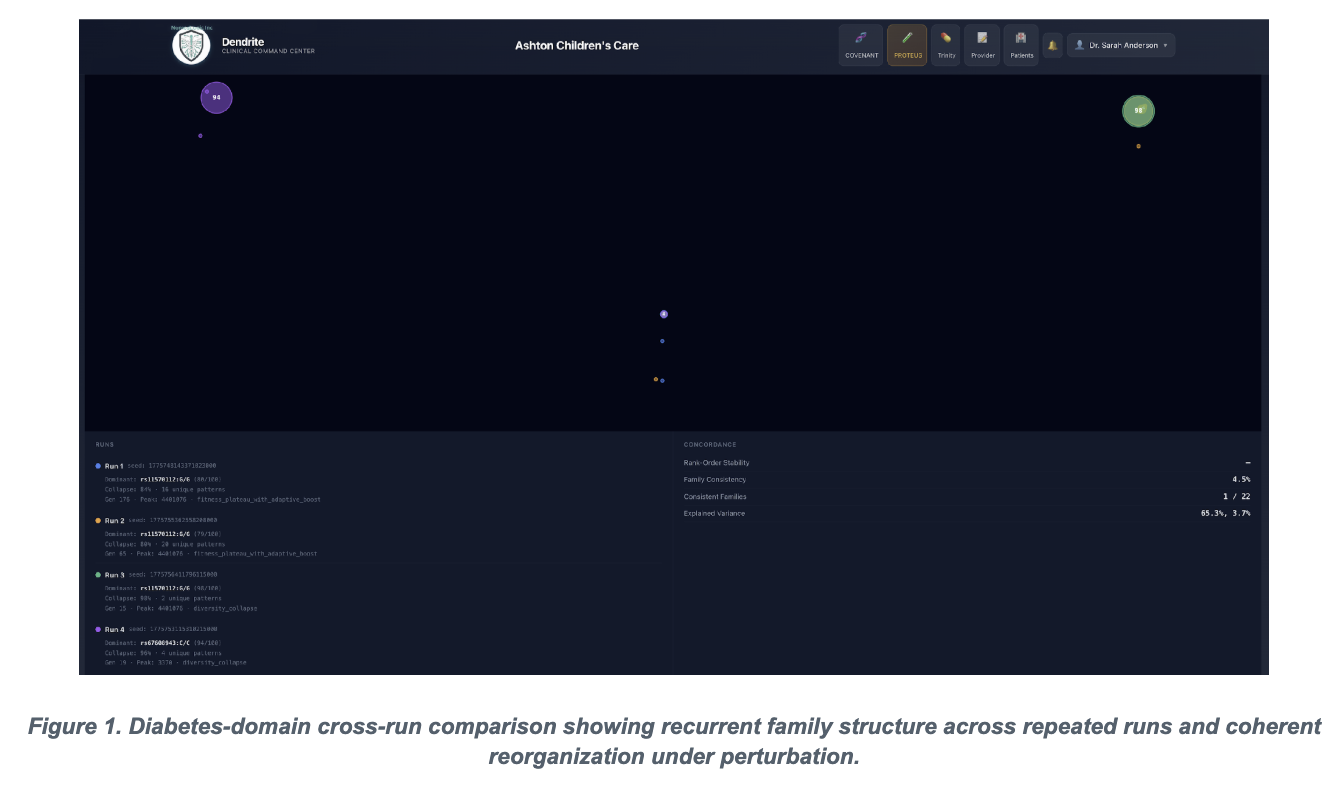
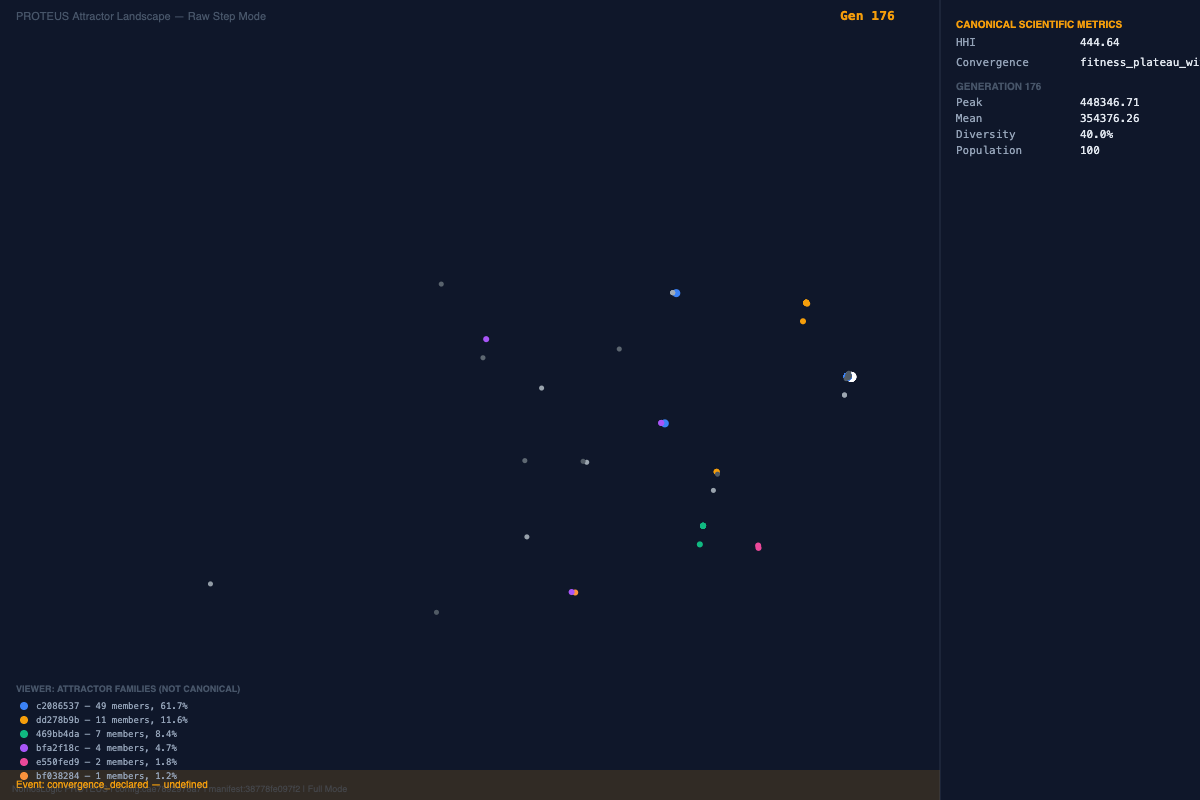
If a genomic system is perturbed and simply collapses, that suggests fragility. But if it reorganizes and returns to a stable structure, that suggests something different: distributed architecture, compensatory behavior, and a deeper systems-level order that cannot be understood by looking at isolated features one at a time.
That is why reproducibility matters so much.
In stochastic or opaque pipelines, it is often difficult to tell whether a result reflects underlying biology, model instability, or procedural noise. Deterministic execution changes that. It allows us to ask a harder and more meaningful question: under identical conditions, does the system consistently resolve toward stable interaction structures?
That is not a cosmetic distinction. It changes what kinds of scientific claims can even be made.
It also changes the downstream implications for drug discovery and precision medicine. If biological systems are more distributed and reconfigurable than our current target models assume, then brittle single-point strategies may fail not because the biology is random, but because the architecture was misunderstood.
The future of genomics will not be built by adding more black-box layers to unstable foundations.
It will be built by treating genomic computation as what it actually is: a systems engineering problem.
And once that shift happens, a different class of infrastructure becomes possible, one built not just to generate outputs, but to produce results that are reproducible, inspectable, and robust enough to matter.