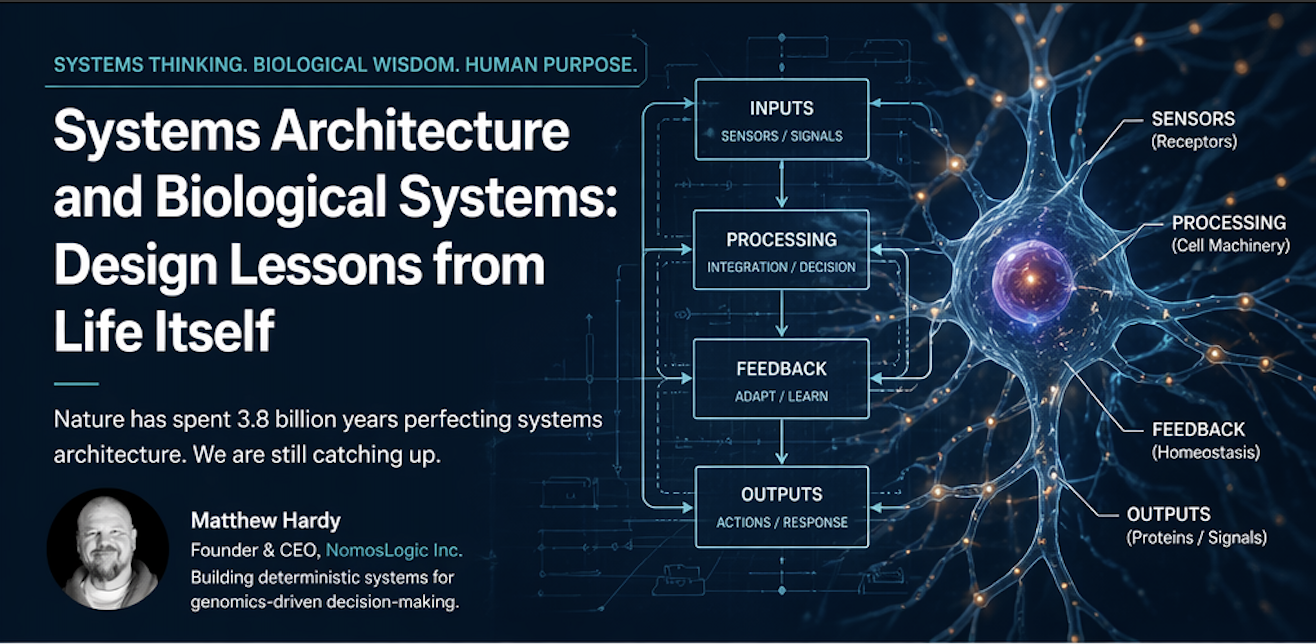
If you’ve ever been told your blood work is "fine" while you still feel sub-optimal, you’ve experienced the failure of population-average reference ranges.
The Bell Curve Trap
Standard lab reference ranges are based on a "Normal Distribution"—essentially the average of everyone else who went to that lab recently (most of whom were sick). At NomosLogic, we reject the idea that a 25-year-old athlete should be measured against the same "normal" range as a 70-year-old sedentary patient.
The NomosLogic Advantage: SNP Integration
The core of our Clinical Logic Objects (CLOs) are the synthesis of static DNA with dynamic blood work.
Static Data: Your SNPs (Single Nucleotide Polymorphisms) tell us your genetic "potential" or "risks."
Dynamic Data: Your blood work tells us your current "reality."
For example, if you have a genetic variant in the MTHFR gene, your "normal" range for Homocysteine is fundamentally different than someone without that variant. By mapping these two data sets, NomosLogic provides a "Biological GPS" that is unique to your specific code.
@nomoslogic - nomoslogic.com